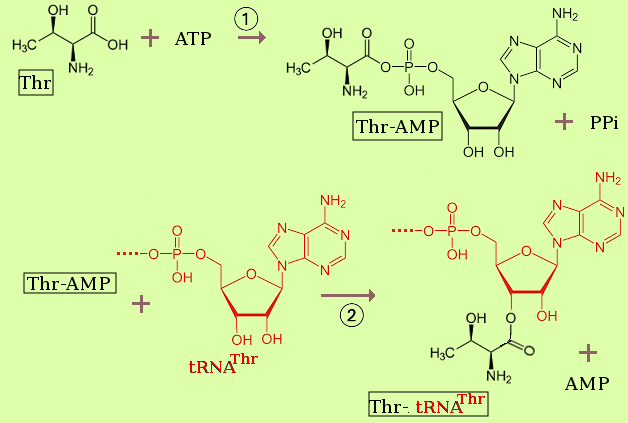
Formation of peptide bonds
There are plenty of Web sites that introduce messnger RNA, ribosomes and other components of the protein biosynthetic apparatus but they sometimes skate over the basic chemistry.
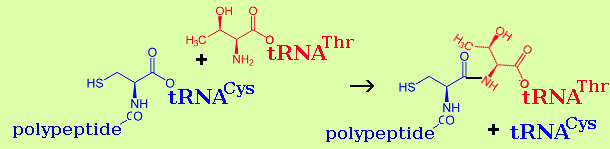
Peptide bonds are formed by an enzyme calatalysed reaction between aminoacyl-tRNA and polypeptidyl-tRNA.
Each of the 21 amino amino acids incorporated into nascent polyppeptid chains has its own transfer RNA ("tRNA") molecules (typically a small family of these). tRNAs are typically of 70-80 nucleotides in length and contain several modified nucleosides.
First we need to attach an amino acid to an appropripate (cognate) tRNA. As an example, threonine is attached by an ester link to the 3'-end of a its tRNA by a reaction that in summary is:

| ester + amine → amide + alcohol |

| polypeptidyl-tRNA + aminoacyl-tRNA → polypeptide+1-tRNA + tRNA |
RNA and ribosomes
J H Parish Undergraduate Lecture on RNA structure, 2000; updated 2009)
A A
(1) 5'....A C A A G A C A C G A C A C G A G
3'....U G U U C U G U G U U G U G C U A
G A
A A
(2) 5'....A C A A G A C A C G A C A C G A G
3'....U G U U C U G U G U U G U G C U A
A A G A
C C
A
G A A A
(3) 5'....A C A A G A C A C G A C A C G A G
3'....U G U U C U G U G U U G U G C U A
A A G A
C C
A
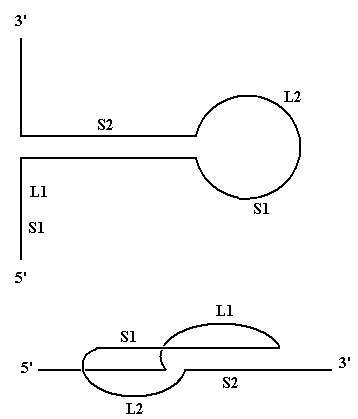
(1) is a stable helix even though it contains a G:U base pair: G:C and A:U
stabilise the helix; G:U has no stabilising or destabilising effect.
(2) and (3) are both less stable but (2) is better than (3) because in (2)
the "upper strand" still has base stacking interactions (between an
A and a G).

A 3'(OH)
C
C
73
(P)5' 1 . 72 acceptor
2 . 71 stem
3 . 70
4 . 69
5 . 68 T--stem-----loop
6 . 67
D--loop---stem 7 . 66 C 59
U 65 64 63 62 61 A
15 A 9 . . . . .
16 13 12 11 10 57
17 . . . . 49 40 51 52 53 C
G T Ψ
G 22 23 24 25 48
20 A 26 47
27 . 43 44 46
28 . 42 45 variable loop
anticodon 29 . 41
stem 30 . 40
31 . 39
32 38
anticodon U 37
loop 34 36
37
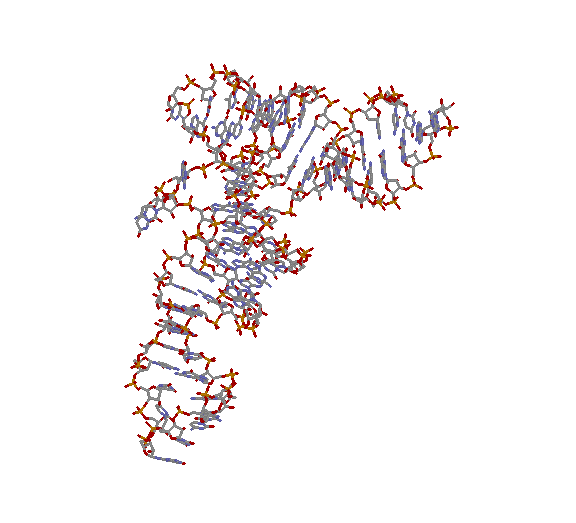


More-or-less a complete issue of Science2000 289: 878...
is devoted to recent developments.
Here is the text of the summary by T R Cech:
The amino acids we obtain by digestion of steak, salmon, or a lettuce salad are loaded onto transfer
RNAs (tRNAs) and rebuilt into proteins in the ribosome, the cell's macromolecular
protein-synthesis factory. The bacterial ribosome is composed of three RNA molecules and
more than 50 proteins. Its key components are so highly conserved among all of Earth's species that a
similar entity must have fueled protein synthesis in the common ancestor of all extant life.
Although the chemical reaction catalyzed by the ribosome is simple--the joining of amino acids
through amide (peptide) linkages--it performs the remarkable task of choosing the amino acids to be
added to the growing polypeptide chain by reading successive messenger RNA (mRNA) codons. On page 905 of this issue, Steitz, Moore, and
colleagues now provide the first atomic-resolution view of the larger of the two subunits of the ribosome. From this structure they
deduce on page 920 that RNA components of the large subunit accomplish the key peptidyl transferase reaction. Thus, ribosomal RNA (rRNA)
does not exist as a framework to organize catalytic proteins. Instead, the proteins are the structural units and they help to organize key ribozyme
(catalytic RNA) elements, an idea long championed by Harry Noller,
Carl Woese, and others.
These landmark publications are but the latest chapter in a progression of ribosome structural studies that have spanned four decades. Early electron
micrographs of ribosomes in action led to immunoelectron microscopy and ultimately to cryo-electron microscopy images of about 20 Å resolution
. Proteins were also located within the ribosome by neutron scattering. However, to achieve atomic resolution, x-ray crystallography
is required, a daunting task given the huge size (2.6 x 106 daltons) and asymmetry of the ribosome. The pioneering crystallization of ribosomes from
the bacterium Haloarcula marismortui in the 1980s by Ada Yonath and H. G. Wittmann provided the first rays of hope, but it is
only in the past few years that crystal structures have been determined for the large subunit (5 Å resolution) (3e small subunit (5.5 Å resolution)
, and the whole ribosome complexed with tRNAs (7.8 Å resolution)
Now, at 2.4 Å, almost the entire chain of the 23S rRNA and its tiny 5S rRNA partner, totaling 3043 nucleotides, have been fitted into the electron
density map of the H. marismortui large ribosomal subunit. The RNA secondary structure (intramolecular base-pairing pattern) of the
large-subunit rRNA had been determined previously , and is present as predicted in the x-ray structure. In addition, a large number of
unpredicted RNA tertiary structure interactions are now seen. Overall, the RNA forms a huge single mass of tightly packed helices, not six discrete
domains connected by floppy linkers as a naïve observer might predict from looking at the secondary structure diagram.
Where, then, are all of the proteins, and what is their function? The globular domains of 26 proteins are found largely on the exterior of the subunit
(see the figure). Twelve of these proteins have unusual snake-like extensions, devoid of tertiary structure and in some cases even secondary
structure, and an additional protein is entirely extended; their shapes are molded by their interactions with the RNA. From these pictures, and from
what is known about protein cofactors that facilitate the action of some other ribozymes, it is likely that these ribosomal proteins buttress, stabilize,
and orient the otherwise floppy RNA into a specific, active structure.
The original contains references and links removed here.
Here is the picture from Cech's article:
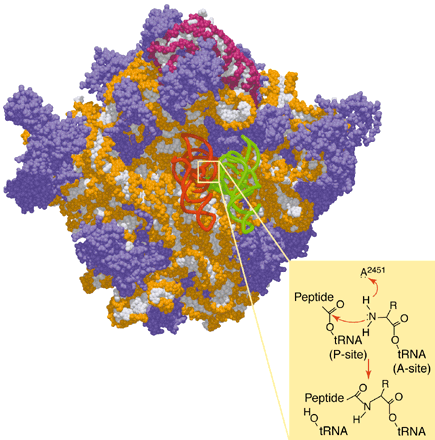
.... and the original text.....
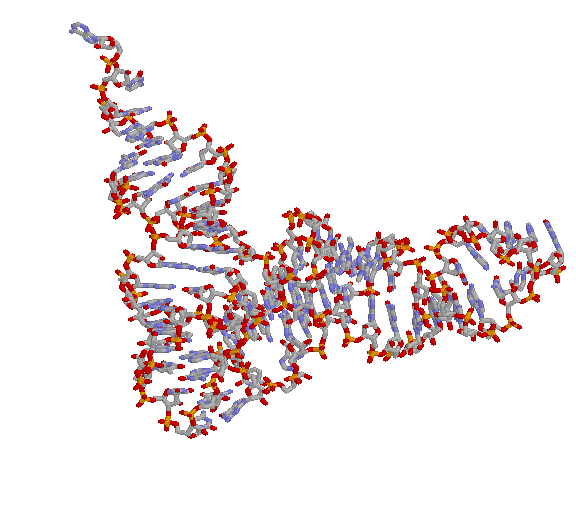
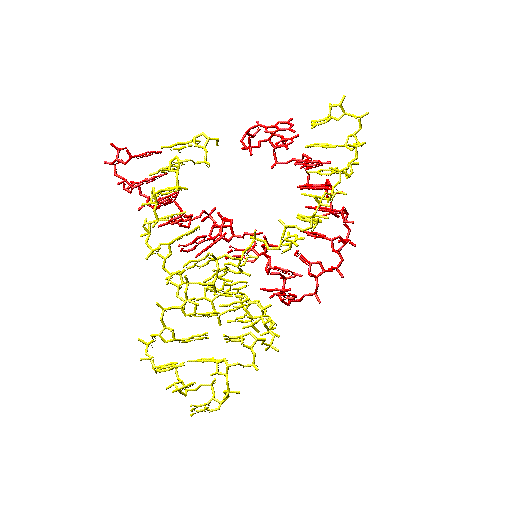
A ribosome's true colors. (Top) The large subunit of the ribosome seen from the viewpoint of the small subunit, with proteins in purple, 23S
rRNA in orange and white, 5S rRNA (at the top) in burgundy and white, and A-site tRNA (green) and P-site tRNA (red) docked according to.
(Bottom) The peptidyl transfer mechanism catalyzed by RNA. The general base (adenine 2451 in Escherichia coli 23S rRNA) is rendered
unusually basic by its environment within the folded structure; it could abstract the proton at any of several steps, one of which is shown here.
Compararable detail for the smaller 30S subunit can be found in the 21st
September 2000
issue of Nature The editorial reads:
The ribosome is the protein factory of the cell, translating the
genetic information encoded in messenger RNA into proteins. In
bacteria, the ribosome is made up of a large and a small subunit
known as the 50S and 30S subunits respectively.
The structure of the small, 30S, ribosomal subunit of the bacteria
Thermus thermophilus is revealed at a resolution of 3 Å
in the journal Nature. In a second article, functional
insights gained from this structure are described, and the
structure of the 30S subunit interacting with antibiotics, which
can interfere with ribosomal decoding and translocation, is
presented.
The ribosome has been intensively studied for the last 40 years
and, as James Williamson says in an accompanying News and
Views article, the structure,
"finally provides a face to players
that have long been known by their functions."
One set of views of the 30S subunit and its legend are below.
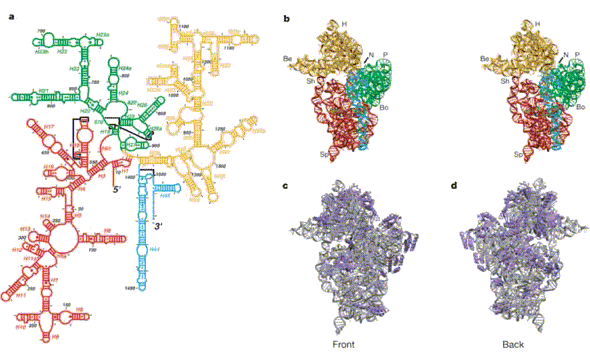

Figure 2. Overview of the 30S structure.
a, Secondary structure diagram of 16S RNA (modified
with permission from
http://www.rna.icmb.utexas.edu/CSl/2STR/Schematics/e.coli16s.27.5.5.schem.ps; see also ref.
21), showing the definition of the various helical elements used throughout the text. The
numbering and diagram correspond to the E. coli sequence. Red, 5' domain; green, central
domain; orange, 3' major domain; cyan, 3' minor domain.
b, Stereo view of the tertiary structure
of 16S RNA from our refined model, showing the 50S or 'front' view, with the same colouring
for the domains. H, head; Be, beak; N, neck; P, platform; Sh, shoulder; Sp, spur; Bo, body.
c, d,
Front (50S) and back sides of the 30S. Grey, RNA; blue, proteins.
Structure and function
animation from MRC Cambridge including excellent animations of the phases of protein biosynthesis.
GOTO TOP
Genetic Code
The phrase "genetic code" has been misused to mean a genome (for example). Rather it is the
relationship of the sequence of mRNA (or the + or non-transcribing strand) of DNA. The code is a non-overlapping triplet (like a language with only 3-letter words). After the general features of the code were established, the code itself was established in the 1960s by groups led by H.G. Khorans and Marshall Nirenberg. They shared a Nobel prize with Robert Holley who was the first to sequence a tRNA molecule and identify an anticodon.
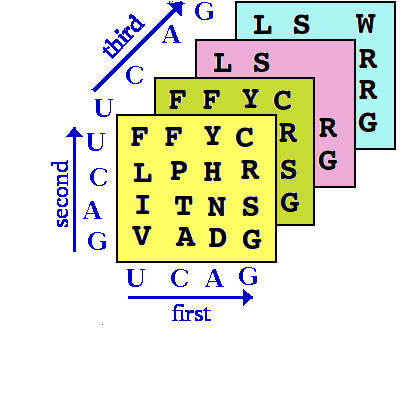
The general nature of the Genetic code is that it is a 3D array which cannot easily be represented on a
flat sheet of paper (or computer screen) but is easily represented as such in computer software (this
speeds up the process of "looking up the code"):
 | U | C | A | G |  |
|---|---|---|---|---|---|
| U | F F L L | S S S S | Y Y ✳ Q in some ciliates ✳ Q in some ciliates | C C ✳ U W (mycoplasmas, mitos...) W | U C A G |
| C |
L T in yeast mitos. L ditto L ditto (below also) L S in C. cyl. |
P P P P | H H Q Q | R R R R |
U C A G |
| A | I I I M in several mitos. M | T T T T | N N K K | S S R ✳,G,S in mitos. R ✳,G,S in mitos. |
U C A G |
| G | V V V V | A A A A | D D E E | G G G G | U C A G |
DNA repair
This lecture was given to the EPSRC CYTOCOM group in 2000.
It is the biological background to a possible computing metaphor. In general if macromolcules become
damaged they are destroyed and re-synthesised. There are four notable exceptions: (1) aminoacyl-tRNA
(if an amino acid is joined to a non-cognate tRNA), (2) the 3'-terminus of tRNA (CCA) which becomes a bit tatty with time, (3) the bacterial cell wall and (4) DNA.
DNA background.
Here is in 1 convention for drawing a DNA double-stranded
molecule. The reason the ends are called 5'- and 3'- is unimportant
(just chemistry) but the fact that they represent directions IS important:
ACAT is different from TACA just as dog is different from god.
5'-...A C A T A G C A T C C A T A G T A C...-3'
The relationship is that the 2 strands are antiparallel (look at the 5' and
3') and the pairing rules are A:T and C:G.
3'-...T G T A T C G T A G G T A T C A T G...-5'
Several things can go wrong: during DNA replication the 2 strands come
apart and the new strands are synthesised 5'-to-3': the enzyme is called
DNA polymerase so here are some steps where the lower strand above is acting
the template. However I've put hyphens in to emphasise that the residues are
all joined up into long chain molecules.
3'-...T-G-T-A-T-C-G-T-A-G-G-T-A-T-C-A-T-G...-5'
5'-...A-C-A-T
The next letter (or 'nucleotide' as we biochemists say) is a A...
3'-...T-G-T-A-T-C-G-T-A-G-G-T-A-T-C-A-T-G...-5'
but now the wrong letter is inserted.....
5'-...A-C-A-T-A
3'-...T-G-T-A-T-C-G-T-A-G-G-T-A-T-C-A-T-G...-5'
DNA polymerase looks at the last 'base pair' it has made and spots it
doesn't fit so the next 2 stages are:
5'-...A-C-A-T-A-A
3'-...T-G-T-A-T-C-G-T-A-G-G-T-A-T-C-A-T-G...-5'
and....
5'-...A-C-A-T-A A
3'-...T-G-T-A-T-C-G-T-A-G-G-T-A-T-C-A-T-G...-5'
so now we can try again and choose (hopefully) a G. The important point
here is that error-checking and correction are parts of the activity of
the polymerase enzyme. There is another biochemical example of this kind
of thing but we'll leave it at that.
5'-...A-C-A-T-A
Watch out you've just had a somatic mutation: As a result of an
enzyme-catalysed reaction, C residues are coverted to U's; we chemists refer
to this an example of a deamination but the important thing is that U behaves
like T:
3'-...T-G-T-A-T-C-G-T-A...-5'
becomes...
5'-...A-C-A-T-A-G-C-A-T...-3'
3'-...T-G-T-A-T-U-G-T-A...-5'
so there is a danger of a mutation... if the upper strand is the template, the
lower one becomes...
5'-...A-C-A-T-A-G-C-A-T...-3'
5'-...A-C-A-T-A-A-C-A-T...-3'
DON'T WORRY ABOUT IT! There is an enzyme constantly scanning your DNA and
marking the U's by putting a cut in the strand which is them repaired by a
method similar to that of the following section but before we look at that
remember DNA does not look like two strings of characters: it is a lumpy
molecule and the repair
enzymes actually detect irregularities in a pseudo-regular structure.
Now here's a different problem. As a result of a chemical attack or UV
or ioinising radiation or simply spontaneously, the DNA becomes damaged....
I've taken the hyphens away to give myself a bit more room. x following a
letter = damaged base.
5'-...A-C-A-T-A-g-c-a C-A-T-A-G-T-A-C...-3'
5'-...A C A T A GxC A T C C A T A G T A C...-3'
There are several pathways for dealing with this but in essense they consist
of 2 steps and there are 2 kinds of alternative.
3'-...T G T A T C G T A G G T A T C A T G...-5'
1 recognition of the damage and
2 doing something about it.
1:
It is the shape of the bad base pair that is recognised (Gx:C in this case).
2:
One route involves simply removing or repairing the damage in situ so an
enzyme whips off the "x". More usually a cut is made by (actually 1 of
2 methods) but for simplicity let's go for the "corrignedase" mechanism.
A corrigendase scans DNA for such bad shapes and would do something
like this... I've stuck the hyphens back in and the x is now over the G.
x
The corrigendase puts a cut* in the region of the damage*.
5'-...A-C-A-T-A-G-C-A-T-C-C-A-T-A-G-T-A-C...-3'
3'-...T-G-T-A-T-C-G-T-A-G-G-T-A-T-C-A-T-G...-5'
*In case you talk to a real biochemist (s)he will point out that there are
usually 2 cuts not quite as close to the damage as I've shown.
5'-...A-C-A-T-A-g T-C-C-A-T-A-G-T-A-C...-3'
Now there is another enzyme that is called a "repair polymeraase" that
concurrently erodes and re-syntheisises the damaged strand.
3'-...T-G-T-A-T-C-G-T-A-G-G-T-A-T-C-A-T-G...-5'
Using little letters to represent repaired strand....
5'-...A-C-A-T-A-g T-C-C-A-T-A-G-T-A-C...-3'
Until the repair polymerase falls off (stochastic process... typically
the repaired bit is around 200 bases not the few shown here) and we
end up with:
3'-...T-G-T-A-T-C-G-T-A-G-G-T-A-T-C-A-T-G...-5'
3'-...T-G-T-A-T-C-G-T-A-G-G-T-A-T-C-A-T-G...-5'
5'-...A-C-A-T-A-g-c-a-r-c-c-a-t-a G-T-A-C...-3'
The last job is to join the a to the G a-G.
3'-...T-G-T-A-T-C-G-T-A-G-G-T-A-T-C-A-T-G...-5'
An interesting feature of this system is that the enzymes "choose" which
strand to regard as the template and which to regard as damaged. Remember
it was the shape of Gx:C that was spotted as bad, how do we say, 'ah
well it is "obvious" that we must use the lower strand as the template in
this case?'. Well the answer is that in reality there are more than 4 bases
in DNA, there are 2 more "called" (well here at least) Am and Cm so our
original molecule (no hyphens) might actually be:
5'-...A C A T A G C A T C C A T A G T AmC...-3'
I've suggested that there are more m's (think of m standing for modified)
in the lower strand. This means that this is the older strand because
only A, C, T, G are incorporated by DNA polymerase and the m's are bodged
in later. So the system has guessed that "our damaged G" is in the less
modified (i.e. "younger") strand. This is correct because it is during
phase of DNA replication that DNA is most sensitive to chemical etc. damage.
3'-...T G T AmT C G T A G G T AmT C AmT G...-5'