 Informatics describes the study and practice of creating, storing, finding, manipulating and sharing information. There are many other neologisms and phrases derived from this: bioinformatics, chemoinformatics, health informatics, nursing informatics, poli-informatics, for example.
Informatics describes the study and practice of creating, storing, finding, manipulating and sharing information. There are many other neologisms and phrases derived from this: bioinformatics, chemoinformatics, health informatics, nursing informatics, poli-informatics, for example.
Bioinformatics is a largely but not exclusively computational subject. In the biological sciences, bioinformatics is so central to understsanding the enormous data sets that characterise modern biology that it is arguably not a separate discipline. Nevertheless the theory of information is extremely important and a comprehensive book can be read on the Web thanks to
David J C MacKay at Cambridge.
The following is the only survivor from the original Web site (this revision August 2015). These notes accompanied a set of lectures on the chemical and biological background to bioiformatics for students who had studied neither chemistry nor biology.
| 1 | Biological chemistry in time and space |
|---|
| 2 | Molecular structures and representations |
|---|
| 3 | Equilibrium and reactions |
|---|
| 4 | The biological and chemical literature |
|---|
| 5 | Macromolecules: primary structures and conformations |
|---|
| 6 | Genomes, gene regulation and protein biosynthesis |
|---|
| 7 | Metabolic, regulatory and neural networks |
|---|
| 8 | Classification and ontologies in biological sciences |
|---|
| 9 | Recombination, repair, rearrangment and evolution |
|---|
| 10 | Exam questions (!) |
|---|
1: Biological chemistry in time and space
| Contents |
Time scales
Size scales
Complexity
|
| Abstract |
The earth has existed for a substantial proportion of the
history of the known universe and has been inhabited for
the greater part of its history.
A useful classification is that of Chandler for the semioitics
of complex systems.
|
| Objectives |
At the end of this topic, you should understand the way in complexity
arose from complex molecules and the interactions of these.
|
| Why is this topic important? |
Because chemistry is important.
|
| Why is it interesting? |
This is a good natural example of emerging properties as a
system becomes more complex.
|
Time scales
Here is a summary of history from the big bang to today.
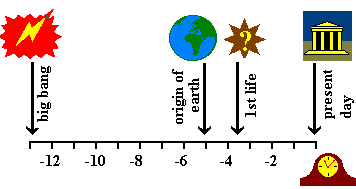
The units on the abscissa are 10-12 years. The scale is
approximate and, arguably, contentious. Also the cartoons might be
misleading: the old world continents were formed quite recently and
we do not know what the first form of life might have looked like.
This form of life
was probably some kind of "bacterium" but the cartoon is
realistic in the sense that the organism was certainly not green. The
point we make is that life on earth occupies a significant proportion
of the "history of everything" on a linear scale on the
abscissa. Although
we look at this in more detail in section 8
we can point out here that a logarithmic scale would be needed to
indicate a separation of "closely related" pairs
of organisms such as codfish & gorillas, carrots & buttercups,
yeast & mushrooms....
GOTO top
Size scales
With the important exception of
radiation biology biological chemistry
and biology are concerned with the scale of small molecules to
whole organisms. In a simple molecule (we take ethylene
[C2H4] a plant hormone, oxygen
[O2] the product of photosynthesis and
methane [CH4] a gas produced by methanogenic archaebacteria
as examples of small molecules of biological origin), bonds between the
atoms are of the order of 1 Å (0.1 nm). This is short for Ångström
unit, i.e. 10-10m, named for Anders Jonas Ångström, a XIX century
Swedish physicist. This is a non-SI unit still
in common use in biological chemistry and molecular biolgy.
Thus the lower end of the scale of size is 1Å.
The sizes of macromolecules are
several orders of magnitude above this.
Viruses are arguably not "forms of life" but the smallest
of these are around 25 nm in diameter. The smallest
"real organisms" are small bacteria, small spheroidal or
sausage-shaped organisms of approximately 10-6m
[1 mu (micron)] and the largest...? there is actually some uncetainty
about the "largest creature on the earth" but the sperm
whale, the giant redwood and the giant squid are all certainly large
so perhaps an upper length is of the order of 102m. As we
have a range of more than 8 orders of magnitude, a logarithmic scale
would be needed to produce a diagram of these sizes.
GOTO TOP
Complexity
Although there is no reason to believe that the "laws of
physics" are mutable, it is clear that as systems become
more complex, new properties emerge. A useful formalisation is
to be found in the semiotics of Jerry Chandler
as summarised in the table below.
|
Symbol |
Class of object |
Notes |
|
Oo1 |
subatomic particles |
electrons, protons, neutrons.... |
|
Oo2 |
Atoms |
|
|
Oo3 |
Molecules |
This class includes ions. |
|
Oo4 |
Biomacromolecules |
DNA, RNA, proteins, polysaccharides |
|
Oo5 |
Cells |
Living objects having a boundary and sustained by a genetic system.
A multicellular organism is 'a cell' in this context. |
|
Oo6 |
Ecoment |
The surrounds of a cell in the above sense.
Nutrients and external signals (such as stimuli) are
parts of the ecoment. |
|
Oo7 |
Environment |
|
Those parts of biochemistry and biology that interface with
bioinformatics are concerned with classes Oo3 to
Oo7 and we highlight such emergent
properties. Next, in topic 2, we shall see the implications of
the 3D properties of molecules and the origins of molecular
asymmetry. In topics 5 and 6 we look at the implications of
the properties that emerge with macromolecules (Oo4)
but here we emphasise three important properties of systems
involving biomacromolecules:
- The very large number of interactions between the component
parts of such molecules result in characteristic shapes
("conformations") which can be of great stability.
- The interactions of certain biomacromolecules (notably proteins)
with smaller molecules result in the specific catalytic properties
of enzymes and also form the basis of signal transduction
(Oo6).
- Interactions between biomacromolecules are also very specific
and form the chemical basis for regulation.
GOTO top
2: Molecular structures and representations
| Contents |
Molecular shapes
Chirality
Conventions
Properties
|
| Abstract |
3D chemical structures of molecules are drawn or described in
one or two dimensions. Important properties of biological
molecules derive from a type of asymmetry referred to as
"chirality"; of particular importance is the
presence of two or more chiral centres in a molecule or in
an interacting molecular system.
|
| Objectives |
In this topic you will learn how to recognise and describe
organic molecules and will appreciate the concepts of conformation,
isomerism and functional group.
|
| Why is this topic important? |
It is the essential foundation for understanding biological chemistry.
|
| Why is it interesting? |
There are several points of interest (I hope) but the ideas behind
representation and interpretation have generic aspects.
|
GOTO TOP
Molecular shapes
We cannot cover all organic chemistry but there is no
shortage of references.
We need to recall the valencies ("combining power") of some of
the elements of importance in biological chemistry.
| Element | Valency | Notes |
|---|
| C | 4 | The chemistry of carbon is called
"organic chemistry" but not all organic compounds
are relevant to biological chemistry. |
| H | 1 | |
| O | 2 | |
| N | 3 | N can have a valency of 5 |
| P | 5 | P can have a valency of 3 |
| S | 2, 4 or 6 | 2 in proteins etc. |
| Fe | 2 or 3 |
usually as +ve ions in biological chemistry |
| Na | 1 |
always as +ve ions in biological chemistry |
| K | 1 |
always as +ve ions in biological chemistry |
| Mg | 2 |
always as +ve ions in biological chemistry |
| Ca | 2 |
always as +ve ions in biological chemistry |
| Cl | 1 |
nearly always as -ve ions in biological chemistry |
We now have enough information to write down the formulae of, for
example, some simple hydrides: methane CH4, water
H2O, ammonia NH3 and also some other simple
molecules such as methanol H3COH and formaldehyde
H2C=O. Note this last case does not violate the valency
rules for C, H and O because of the bonding arrangements. There are
four bonds in CH4, each consisting a shared pair of
electrons, whereas in H2C=O, there is a double bond
(i.e. 2 such shared pairs) between the C and the O.
What are the shapes of some simple molecules? Some are shown in the
diagram.
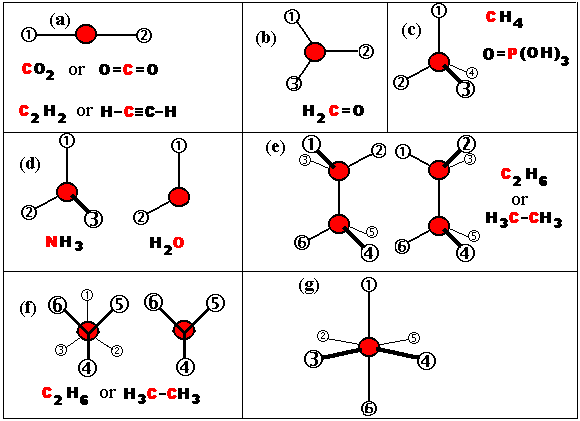
Some molecues have the atoms in a straight line (a). In others the
molecule is flat (b), i.e. the four atoms of H2C=O all lie
in one plane. In others (c) a central atom is surrounded by four
others that form the vertices of a tetrahedron. The reasons for these
shapes are well understood but here we are
concenrned only with the shapes and we emphasise the tetrahedron is
not found only in carbon compounds: the other example in (c) is
phosporic acid. (d) illustrates NH3 and H2O;
the former is shaped like a 3-spoked umbrella cover and water is
"bent": both are based on the tetrahedron but with one
(in the case of NH3) or two (in the case of H2O)
vertices missing. Before leaving the tetrahedron, ethane (e)
illustrates the fact that rotation is allowed on the axis of a C-C
bond and the two pictures are two extreme conformations of the
molecule. (f) illustrates these two confomations from a differrent
viewpoint. Another shape found in biological molecules, notably
around complex metal ions is the octahedron (g).
GOTO TOP
Chirality
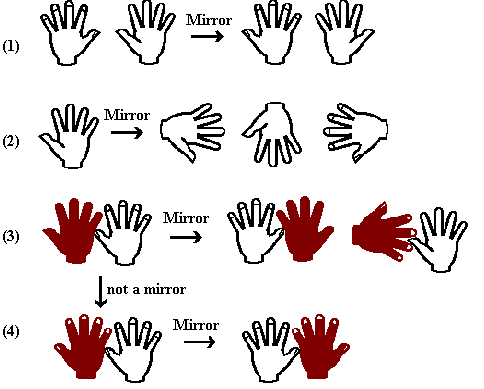
Chirality is the chemical name for 'handedness'. The picture
below revises handedness: (1) a left and right hand and the mirror
image. (2) The mirror image of a left hand is a right hand: note
that by "mirror image" we mean that the right hand is
the "left hand in a looking glass world", i.e. we are not
concerned about the orientation of the hands. (3) a left-right
pair produces a right-left mirror image. The transition from
(3) to (4) was achieved by taking a mirror image of the dark hand
and leaving the white one alone. This is a new pair but of course
has its own mirror image.
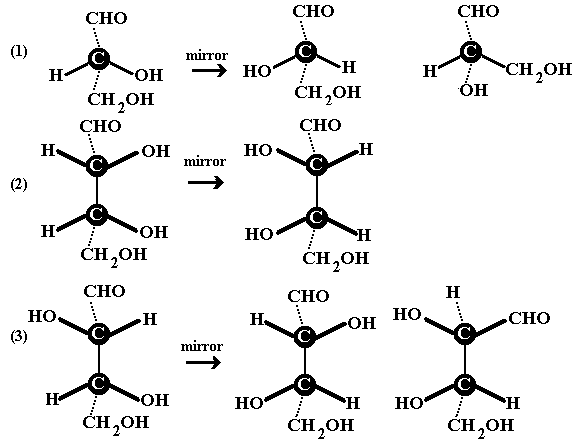
Below we look at equivalent pictures for some molecules. (1) is
molecule with a chiral centre, i.e. a tetrahedral C atom
with 4 different substituents. (2) is a molecule with two such
centres. (2) and (3) are related in the same ways as the pairs of
hands in (3) and (4) in the sketch above. Note that not only
do we ignore the orientation of the "looking glass"
structure but we are not concerned with the
conformations.
GOTO TOP
Conventions
Organic chemistry has a strict set of rules for
naming substances. However many biochemists and most molecular
biologists have a cavalier approach to naming things. Here we
are even more irresponsible and just give things names if we need
to refer to them later. However we can look at methods for
representing 3D structures in 2D. Let's make up some rules:
| Objects | Viewpoint/conformation |
Conventions |
|---|
| hands |
from the back of the hand |
draw arrow from wrist to fingers and draw line for the thumb |
| Certain molecules |
Look at each chiral C atom so that attached C atoms are
away from the viewpoint and the other two are towards the viewpoint;
use a conformation so this applies to all chiral C atoms. |
Represent C-C bonds just as a line;
do not bother to draw C-H bonds. |
Here are the applications of the rules:
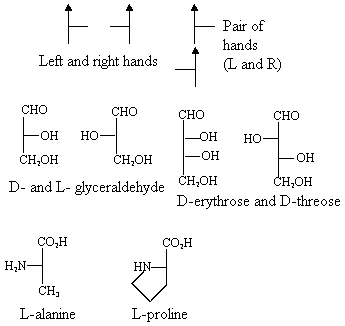
The 2nd line of the picture corresponds to some of the molecules we
saw earlier. We have given them some names also. Note that
D-glyceraldehyde is a very simple sugar. L-alanine and
L-proline are examples of amino acids, the building blocks of proteins.
Two molecules such as D- and L- glyceraldehyde are called
enantiomers and two such as threose and erythrose are called
diastereoisomers.
The letters, D and L, are derived from the characteristic property of
enantiomers: in solution they rotate the plane of plane-polarised
light and are said to be optically active. This is the only
property that differentiates such enantiomers unless they interact
with other enatiomeric substances. Originally the letters were
written in lower case and stood for dextro (right)
and laevo (left). However there is no intrinsic
significance in the d/l value and in biological
chemistry the absolute configuration of the chiral C atom is important.
All amino acids in proteins have the "L" configuration and
likewise most carbohydrates, of which sugars form a sub-set, are
"D". There are chemical rules for
labelling such chiral centres but these have limited use in
biological chemistry.
A "rule" (more exactly a mnemonic) used by biochemists for
describing the absolute configuration of an L-amino acid is
"CORN".
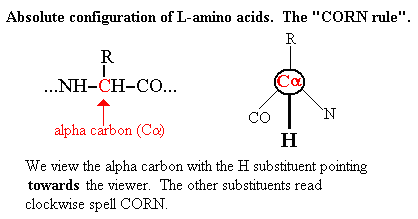
However the advent of computational chemistry and chemo- and
bio- informatics has led to a requirement for a 1D or string
representations of structures including chirality. The
system is called
SMILES.
Examples of SMILES strings are N[C@@H](C)C(=O)O for L-alanine and
N1[C@@H](C)CCC1C(=O)O for L-proline (this latter example shows how
SMILES can accommodate cyclic structures).
A way to look up SMILES strings, is to use the
CADD W3
site.
Before we leave this point, we can understand how a SMILES string
(or name) can be used to describe a structure that can be drawn
as sticks etc. However it is important to remember that all these
representations are abstractions. Let us take another look at
L-alanine. Here we know the actual co-oridinates of the atoms in
at least one actual conformation.
In these pictures, C is dark grey, H pale grey, O blue and N red.
Both show the same view of the molecule. On the left is a "ball
and stick" model and on the right a space filling model. To gain
an impression of the structures of biological molecules, visit one of
several Web sites of which
www.nyu.edu/pages/mathmol/library is an
excellent example.
GOTO TOP
Properties
The representation of a double bond by "=" is misleading
because the two "bonds" are not equivalent. There is an
axial shared electron pair and this is surrounded by
π (pi) electrons. However an important
difference between single and double bonds is that latter are
not axes of free rotation so that (picture below) the two
molecules in (a) are different substances: this is an example of
geometric isomerisms and note the naming of the isomers
as cis and trans. In the case of single bond
(b) the isomerism does not occur because these are merely different
conformations. Geometric isomerism is not restricted to double-bonded
molecules: (c) illustrates two geometric in a cyclic molecule. One of these
(trans) is also drawn in a convention that is found in several
(bio)chemical books.
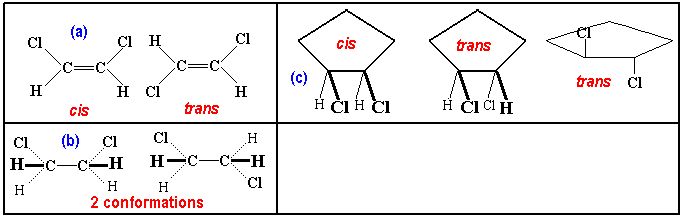
The next picture illustrates some chemcial properties associated with
certain functional groups found in organic molecules.
We introduce two more bits of shorthand: Me- and Et- are
abbreviations for CH3- (methyl) and
CH3CH2- (ethyl) respectively.
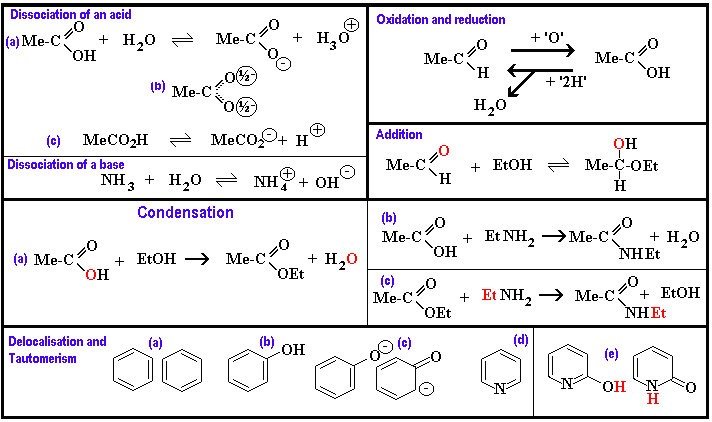
Dissociation of acids and bases: The group -CO2H
is the "carboxylic acid" group. Our carboxylic acid is
acetic acid and in water it forms an equilibrium mixture with the
-vely charged cognate anion (acetate) and a hydrated hydrogen
ion (correctly the "oxonion ion") H3O+
as shown in (a). (b) emphasises that the two C-O bonds in the acetate
ion are not different as (a) implies: the
π electrons are delocalised so each O
atom carries half a -ve charge. Note it is this delocalisation
and the consequent spreading of the load of carrying a negative
charge between 2 O atoms that means that acetic acid can dissociate
and "is an acid" unlike (for example), ethanol Et-OH. (c)
shows the conventional way of writing the equilibrium: we take it
as read that water has reacted and that H+ is in reality
H3O+ and not a proton (sub-atomic
particle).
Ammonia is a weak base, i.e. it reacts with water and in the
equilibrium mixture OH- (hydroxide ions) are formed.
Oxidation and reduction: In the illustration (acetaldehyde and
acetic acid) "oxidation" is the addition of O (or the
abstraction of 2H) and "reduction" is the addition of 2H
(or the abstraction of O).
Addition: A simple reaction of the form:
A + B → C
is an addition. The example shown would seem to be obscure
to a reader of elementary chemistry texts but is important in
carbohydrate chemistry. The red O is
to clarify which parts of the addition product come from where.
Condensation: A simple reaction of the form:
A + B →
C + D
where D is a simple molecule such as water is a
condensation. Again red
atoms are coloured to clarify the
reactions:
(a) is the reaction of an acid and an alcohol to form an
ester
(b) is the reaction of an acid and an amine to form an
amide.
both of these reactions could, in principle, be reversed by the
reaction of the ester or amide with water and these reactions would be
examples of hydrolysis.
(c) is another method of forming an amide (it is the method used in
protein bioisynthesis) and in this an alcohol not water is removed.
Delocalisation and tautomerism: we saw that the
π electrons in the acetate ion are
delocalised. Other examples are benzene (a) and its
derivatives. The two structures are referred to as the
canonical forms of benzene.
(b) phenol, hydroxybenzene. Phenol is a weak acid because
the negative charge is delocalised: two canonical forms of the
phenate anion are shown in (c).
(d) is pyridine, i.e. it is similar to benzene except
one of the CH groups in the ring has been repalced by N.
(e) What are these?... "hydroxy-pyridine" or something
else (it would be called "pyridone")? Answer:
yes !. These are not "canonical forms"
because we have moved, not just π
electrons but an H atom as well but the fact remains that the two
structures in (e) are in equilibrium, they cannot be separated
and are referred to as tautomers.
GOTO TOP
References
Apart from links on the Web (
freebookcentre.net/Chemistry/Organic-Chemistry-Books.html), a complete newcomer to organic
chemistry might wish to read (or even buy) a textbook. The simplest
advice is to browse in a library or bookshop and find a book that
starts from where you want to start. A short and reasonably
inexpensive solution is:
Patrick G.L. (2000) Instant Notes in Organic Chemistry
Oxford: BIOS Scientific Publications.
Of the many big fat texts, I like the English edition of
"Beyer/Walter":
Beyer H., Walter W. & Lloyd D. (translator and editor) (1997)
Organic Chemistry. Chichester: Albion Publishing.
3: Equilibrium and reactions
| Contents |
Units
Equilibrium
Thermodynamic considerations
Reaction rates |
Abstract |
Equlibrium positions are related to classical
chemical thermodynamics. Chemical reactions
need to be thermodynamically feasible and there
needs to be an available kinetic mechanism for
them to occur.
|
| Objectives |
At the end of this topic you should understand
equilbrium constants, pH, pK,
enthalpy- entropy- and free energy- changes,
and rates of reaction.
|
| Why is this topic important? |
It is central to knowing whether reactions can take place.
|
| Why is it interesting? |
Because it is not too far away from physics.
|
GOTO TOP
Units
| Unit | Explanation | Notes/examples |
|---|
| Dalton |
unit of relative molecular mass (RMM) |
We do not here enter the controversy over the use of the
term "molecular weight".
The atoms have relative atomic masses, examples (approximately)
are H 1, C 12, N 14, O 16 and thus the RMMs of H2O and
CH3CH=O are 18 and 44 respectively. |
|
mol |
Gram molecule; the RMM in grams |
1 mol of water is 18 g;
1 mol of CH3CH=O is 44 g. |
| M |
molarity (M stands for molar);
a concentration in mol l-1. |
Note the SI prefixes p, n, m
... can all be used with mol and M. |
| [ ] |
concentration |
[X] means "concentration of X" (e.g. in M)
|
| K or K, Keq etc. |
equilibrium constant |
Given a reaction R1 + R2 ... ↔ P1 + P2 ...
K = ([P1][P2]...)/([R1][R2]...)
|
| k |
rate constant |
for the reaction
R1 + R2 ....→ products
The rate of reaction = k[R1][R2]... |
| p | pX = log10(1/X) |
NB: "p[H+]" is written pH
|
GOTO TOP
Equilibrium
From the table, we can see that for a reaction
such as:
X ↔ Y + Z
in which substance X dissociates to form Y and Z, theres will
be an equilibrium constant, K:
K = [X]/([Y].[Z])
and, as K is a constant (at a given temperature etc.), the
equation can be used for calculating the value of two of the
concentrations, given a knowledge of the other one.
The dissociation of acetic acid in water
is an example of such an equilibrium. If we call acetic acid, AcOH,
the equation becomes:
KA =
([H+].[AcO-)/[AcOH]
(KA is the acid dissociation constant). Likewise
the solution of ammonia in water can be represented as
NH3 + H2O
↔ NH4+
+ OH-
KB =
([NH4+].[OH-])/[NH3]
Don't worry about the seeming loss of [H2O] .... water
has an effective unit acitivity in this reaction... in other
words, "if an H2O molecule is needed, there'll be one
on hand".
Acetic acid is a "weak acid" and ammonia is a
"weak base" which is another way of saying that the two
constants KA and KB are very
small. We take as examples a strong acid, HCl and a strong base NaOH.
In solution these are effectivley completely dissociated:
HCl
→
H+ + Cl-
NaOH →
Na+ + OH-
Thus in 1mM HCl [H+] is 10-3 and the pH is 3.
What about 1mM NaOH? For this we need to know K for the
dissociation of water itself... it is 10-7 and thus the
pH of pure water is 7 (referred to as neutrality) so the pH of
1mM NaOH is 10.
Now we'll have another look at the equation
KA =
([H+].[AcO-]/[AcOH]
Rearrange and take logs to the base 10 (remembering what we mean by
"p") and you should end up with:
pH = pKA +
log10([OAc-]/[HOAC])
Now we look at what happens if we have a mixture of HOAC and a salt
such as sodium acetate (NaOAc) both in solution. To first,
approximations, [AcO-] is the same as [NaOAc] because it
is a salt and dissociated and [HOAc] is the conentration of acetic
acid we put in the solution (provided, as is the case with acetic
acid, KA is very small. So for such solutions
containing a weak acid and a salt of that acid, the equation is
one beloved of biochemists who call it the Henderson-Hasselbalch
equation:
pH = pK +
log10([salt]/[acid])
Clearly this has some applications, given the pK of acetic acid
(it's around 4.0) and we are asked to go into a chemistry lab and
make up a solution of, say, pH 5.1, we can use the equation to work
out how much HOAc and NaOAC to weigh out and dissolve in water.
However the importance of this equation is that the solution is an
example of a buffer. This refers to the fact that the
pH is relatively insensitive to addition of small amounts of acid and
alkali. It is left as an exercise for the reader
to convince herself(himself) of the justification for this claim.
The important point is that biolological fluids whether within cells,
in fluids such as blood or in a solution for experiments in a
biochemistry are all buffered with mixtures (not typically the
acetic acid/acetate system though).
GOTO TOP
Equlibrium
As this module is designed for people with a physics/maths background
I shall simply remind you of some thermodynamic terms but note that
(bio)chemists tend to use Gibbs rather than Helmholz free energies.
[There is a bit more help as an appendix under the "Mol. Biol." tab.]
| Symbol | Meaning | Equation/explanation |
|---|
T
R | Absolute temperature
Universal Gas Constant |
|
| H | Enthalpy or "Heat Content" |
H = U + pV
where U is internal energy; p, V are
pressure and volume. |
| S | Entropy |
A measure of disorder in a system |
| F (or A) | Helmholz Free Energy |
F = U - TS |
| G | Gibbs Free Energy |
G = H - TS |
| a | Activity |
a thermodynamic function that describes the effective concentration
of a substance; for many cases in solution, for substance X,
aX is approximately [X]. |
| Go | Standard free energy |
G at unit activity; it appears as a constant
in equations such as the following which describes G for
the ith component of a system:
Gi = Gio +
RTlnai
|
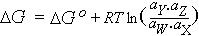 |
Standard free energy change |
Take the reaction W + X ↔ Y + Z
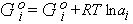
i.e. RTlnK IFF we accept the
"a is [ ]" approximation.
|
| ΔG |
Free energy change |
For the reaction W + X → Y + Z
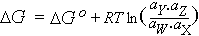
|
|
|
The importance of these variables and formulae in chemistry is
the free energy change) ΔG
must be negative
(i.e. free energy is
"lost" from the system for a reaction to be
thermodynamically feasible). Note that the standard free
energy change
can be calculated from the
equilibrium constant (K) and such values are tabulated but
they are meaningless without a a knowledge of concentrations (the
free energy change in a system at equilibrium is necessarily zero).
A system can however escape from the contraints of the maxim
"Free energy change must be negative"
If the reactions are coupled:
- A + B → C + D
- C + E → F + G
- F → H + J
.... the overall free energy change can be -ve even if one or more
reactions
(say 1) is non-feasible (+ve free energy change) provide the arithmetic
sum of the ΔGs is negative. Thus (a) very
feasible or
favourable reaction(s) can "provide the free energy"
to drive an unfavourable reaction in the chain. This is very important
in biochemistry because there are many highly unfavourable reactions
such as:-
pumping of nutrient molecules up a concentration gradient
-
formation of bonds in the synthesis of complex compounds from
simple ones
-
biosynthesis of nucleic acids and proteins*
*This last example is doubly unfavourable: not only is
bond formation "difficult" but we are creating order out
of chaos: at a point in protein biosynthesis, out of a total of
20 possible amino acids to choose one, we select just one.
That is a highly unfavourable entropy change.
GOTO TOP
Reaction rates
A negative free energy change is a necessary but not sufficient
condition for a reaction to take place. There has to be
an available mechanism for the reaction and the reaction should
occur at a reasonable rate. For example the equlibrium position
for the intercoversion of diamond and graphite lies in favour of
graphite at everyday temperatures and pressures. However the
mechanistic and kinetic considerations do not allow the reaction to
take place so that owners of diamond rings and brooches do not need
to worry about their jewels turning into soot.
Rate constants
A rate constant, k appears in differential equations that
describe the rate of reaction. For example in a simple reaction
A→B
let a be [A] at time 0 and at time t,
[A]=(a-x) then
rate of reaction = dx/dt = k(a-x)
If we integrate this, evaluate the constant of integration by
noting x=0 when t=0 we have:
kt = ln(a/(a-x))
For a slightly more complicated case, A +
B→C
and using the same conventions,
dx/dt = k(a-x)(b-x)
which can also be integrated.
What about "really complicated reactions"? .... in fact
reactions of the sort written down as "balanced equations"
in school chemistry texts are all broken down into simple reactions
with one reactant (unimolecular) or two reactants
(bimolecular). Genuine trimolecular reactions are chemical
curiosities.
What does k mean? Let us take the simple reaction molecules
X and YZ. The reaction is bimolecular and the products are going to
be XY and Z.
 The state represented by
The state represented by
X――Y――Z is a
transition state and represents an energy peak. Popular
and/or elementary chemistry texts are keen on producing a graph,
sometimes unhappily lacking axes, of
this process. However the important point is that Arrhenius described
the relationship between k and E, the activation
energy:
k = Ae-E/RT
We pause here to note that as E is often an order of
magnitude greater than RT, k is very sensitive
to temperature and hence E and associated thermodynamic
parameters of activation are accessible to measurement.
Arrhenius thus made the first breakthrough in relating reaction rates
to molecular processes. Hinshelwood made the second: A in
the Arrhenius equation is not a constant but a probalistic term.
Going back to XY and
Z it is important in most cases that
the molecules are the right way round in 3D for the transition
state to form. This is an entropy effect and if we follow
Hinshelwood and replace A by e-S/R
the equation becomes
k
= e-S/Re-E/RT
= e-(E-TS)/RT
Taking logs....
-RTlnk = E-TS
(-RTlnk) is the free energy of activation
E and S are the energy of activation and
entropy of activation
GOTO TOP
4: The biological and chemical literature
| Abstract |
Use the Web
|
| Objectives |
In this topic, you will be introduced to important W3 sites and
search engines.
|
| Why is this topic important? |
We need to get some more data and ideas.
|
| Why is it interesting? |
Because some of the Web sites are interesting.
|
| Contents |
There are no subsections in this topic.
|
The chemical and biological literature is the primary authorative
resource. Here we concentrate on introductions and tutorials.
It is with some caution that one recommends a lot of W3 sites because
people keep changing their URLs so here is a minimum list.
| URL | comment | more comments |
|---|
| www.google.co.uk |
Useful search engine |
String some words together: it is useful to include
"tutorial" and/or "intro" in the search field.
|
| www.chemfinder.com |
Fairly friendly way of asking "what is that compound?" |
with non-standard names it is better with medical/pharmaceuticals |
| www.ncbi.nlm.nih.gov |
The most important site for molecular biology etc.: you must
get used to it. |
To get started choose their link to "seven modules" |
| www.cas.org |
"cas" is Chemical Abstracts |
There is a different approach from www.ncbi.nlm.nih.gov but there is
considerable overlap in coverage. Try the "SciFinder" and the
life sciences links. |
| www.expasy.ch |
Major Swiss molecular biology server |
largely a research facility but with tutorial material |
| www.expasy.ch |
Introduction to Expasy |
see above |
The remaining topics in this module do NOT contain references
as you should have no difficulty in finding the relevant material
from now on.
GOTO TOP
5: Macromolecules: primary structures and conformations
| Contents |
Amino acids and proteins
Carbohydrates, RNA, DNA
Protein structure
DNA structure
|
| Abstract |
Proteins are polypeptides. Polysaccharides contain
sugar residues. RNA and DNA are polynucleotides.
|
| Objectives |
At the end of this topic you should have a basic knowledge
of the ways in which proteins and nucleic acids are built
up from amino acid and nucleotide residues and ways in which
protein and DNA structures can be represented.
|
| Why is this topic important? |
It underlies most of the applications of bioinformatics.
|
| Why is it interesting? |
It is a classic example of emerging properties.
|
GOTO top
Amino acids peptides and proteins
Proteins are polymers consisting of chains of amino acids:
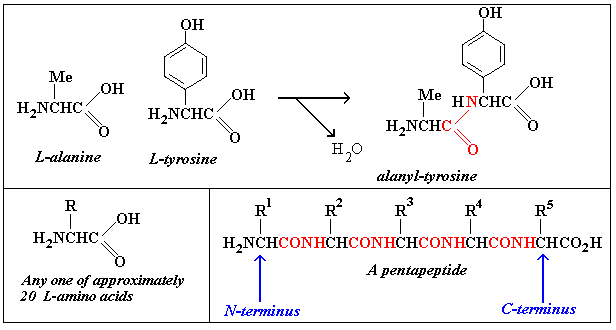
In the picture, L-tyrosine is one of 3 amino acids that contain
an aromatic group, i.e. one that contains a benzene ring.
It is important to note that in this topic we show some apparent
condensation reactions but in no case does this reaction with the
seeming production of water occur during the biosynthesis of the
polymer. The picture introduces a number of conventions and
abbreviations. Peptide bonds are drawn in
red. Small peptides such as the pentapeptide are often just
called "peptides": proteins are polypeptides
with, typically some hundreds of amino acids.
In the table we summarise the 20 amino acids that are most commonly
introduced into proteins during their biosynthesis. The first column
is the one-letter abbreviation usually used in bioinformatics; the
other columns are the 3-letter abbreviations sometimes used in
biochemistry followed by the names. The last column in
this table gives some indication
of the chemical proerties of the R-group. Note that in glycine,
the R group is H so it is neither D- nor L-.
A good representation of 20 amino acids is on a "blogspot".
| A | Ala | L-alanine | small, unreactive |
|---|
| C | Cys | L-cysteine | contains SH |
|---|
| D | Asp | L-aspartic acid | carboxylic acid |
|---|
| E | Glu | L-glutamic acid | carboxylic acid |
|---|
| F | Phe | L-phenylalanine | bulky, aromatic, hydrophobic |
|---|
| G | Gly | glycine | very small |
|---|
| H | His | L-histidine | reactive, can be acid or base |
|---|
| I | Ile | L-isoleucine | bulky, hydrophobic |
|---|
| K | Lys | L-lysine | basic |
|---|
| L | Leu | L-leucine | similiar to I (Ile) |
|---|
| M | Met | L-methionine | contains S, hydrophobic |
|---|
| N | Asn | L-asparagine | an amide |
|---|
| P | Pro | L-proline | an imino acid;
breaks an H-element (except in certain integral membrane proteins) |
|---|
| Q | Gln | L-glutamine | an amide |
|---|
| R | Arg | L-arginine | stronly basic |
|---|
| S | Ser | L-serine | contains OH |
|---|
| T | Thr | L-threonine | contains OH |
|---|
| V | Val | L-valine | very similiar to I (Ile) |
|---|
| W | Trp | L-tryptophan | bulky, aromatic, hydrophobic |
|---|
| Y | Tyr | L-tyrosine | a phenol |
|---|
GOTO top
Carbohydrates, nucleotides, polynucleotides
The picture below illustrates the way in which glycosides are formed.
A simple sugar such as glucose exists in solution as a mixture of
two isomers which are formed by an addition reaction. Solid
crystalline glucose is pure alpha but if it is dissolved
in water it forms slowly an equilibrium mixture of alpha and beta.
The O present in a C=O bond in the straight
chain appears a C-OH in the cyclic forms and
this is referred to as the anomeric OH group.
Below that we show ribose and one of the
products of such an addition reaction.
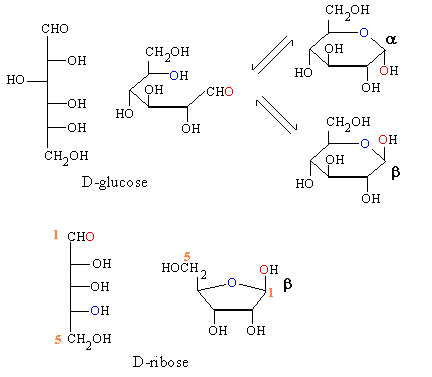
These cyclic forms of sugars such as glucose and ribose are in
solution in equlibrium with other forms but if a condensation occurs
between the anomeric OH and an alcohol (or amine) the products are
glycosides and these are not parts of "alpha/beta
equilibria".
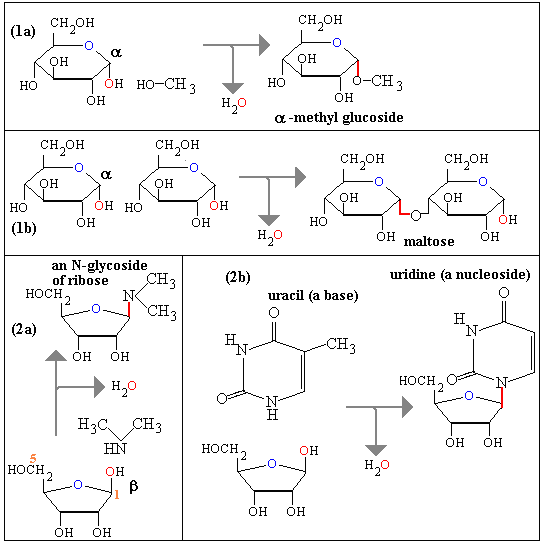
In the following picture we see how the
condensation of the anomeric OH of glucose with an alcohol leads to
a O-glycoside: the examples are a methylglycoside (1a) and
maltose (1b). In contrast if the alcohol is replaced by an amine we get an
N-glycoside. There are two examples here, both N-glycosides
of ribose: (2a) is an illustration with dimethylamine and (2b) shows an example
where the amine is a nucleic acid base and the resulting N-glycoside
is an example of a nucleoside. Glycoside bonds are coloured
red.
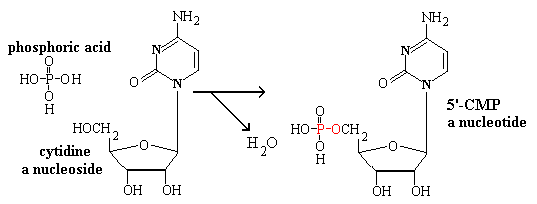
The phosphate esters of nucleosides are called nucleotides.
Polynucleotides (RNA or DNA) phosphopdiesters formed as
follows. RNA molecules contain hundreds or thousands of nucleotide
residues; DNA molecules can contain millions. RNA and DNA differ
in the nature of their sugar component. The blue
OH group in RNA is replaced by H in
DNA. The link is formed by the consensation of a nucleoside triphosphate with the 3'-end of a
growing chain with the elimination of pyrophosphate.
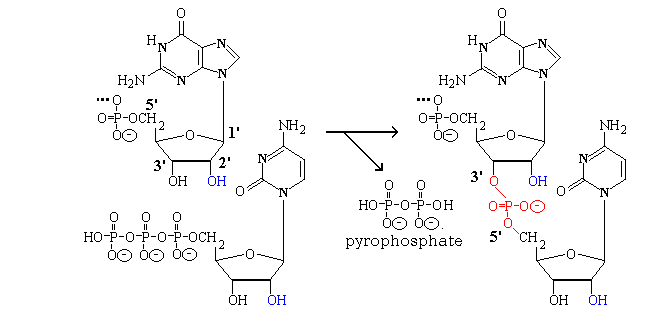
Note that the numbers of the atoms in the ribose (deoxyribose
in DNA) are written 1' ("one primed") etc. to distinguish
them from numbers one might use for the bases. The
phosphate joins a 5'- to a 3'- O. Note
also that we draw phospate as ionised. This is because the pK for
this group is very low and in vivo RNA and DNA are present
as polyanions: the counter ions are metals and/or positively
charged (basic) side chains in proteins. Thus although DNA and
RNA stand for (deoxy)ribonucleic "acids", they should really
be called (deoxy)ribonucleates. The usual convention
for drawing a nucleic acid (a fragment of RNA in this case) is:
5'...GC 3' in the picture,
5'-to-3' is the equivalent of N-to-C in a polypeptide. Again,
5'-to-3' is be default left-to-right but sometimes it is useful
to draw the sequence backwards in which case the labelling must be
explicit:
3' CG...5'
The abbreviations etc. for the components of RNA and DNA are summarised
in the following table.
| | | type | comp. residue |
|---|
| U | RNA | pyrimidine | A |
|---|
| T | DNA | pyrimidine | A |
|---|
| C | DNA and RNA | pyrimidine | G |
|---|
| A | DNA and RNA | purine |
T (U in RNA) |
|---|
| G | DNA and RNA | purine |
C |
|---|
| Y | DNA and RNA | any pYrimidine |
R |
|---|
| R | DNA and RNA | any puRine |
Y |
|---|
We bother here neither with the structures etc. nor the names of the
nucleosides and nucleotides. The last two rows in the table indicate
abbreviations and these are useful in summarising one property of
the "complem. residue". DNA and RNA molecules contain
double helical secondary structures that we can leave for the moment
except to point out that they involve the pairing of complementary
pairs so that part of a double stranded DNA molecule might look
as follows.
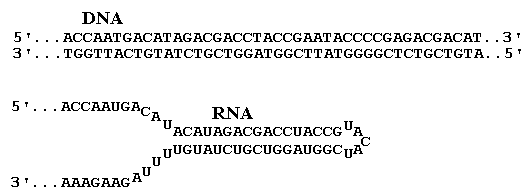
RNA molecules (with some viral exceptions) do not consist of two
separate strands but the same rules of complementarity apply to
short intramolecular stretches of complementarity.
GOTO top
Protein structure
A bioinformatic understanding and view of protein structure and the
ways in which such structures are classified are covered
elsewhere but here we summarise a few points
and methods of representing such structures. We choose just one
example, a protein called pCro. At this stage it does not matter
too much what pCro is for but it has certain advantages, notably
it is very small and easy to represent. There follow 3 sketches
of pCro, all drawn from the same viewpoint.
First a "ball and stick" model: the colour coding
is C grey, O red, N blue and S yellow.
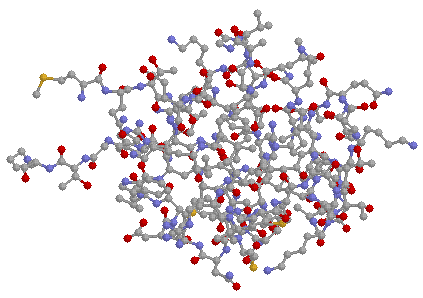
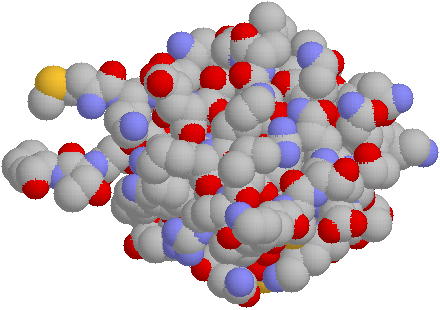
Next a space filling model: if does not tell us much except we can
see one of two S atoms poking out on the surface and it emphasises that
the structure is very compact.
Finally a cartoon of pCro drawn to illustrate the
secondary structure elelments of the structure. In this
case most of the structure is made up of alpha-helix (often
simply "H" for helix in bioinformatics).
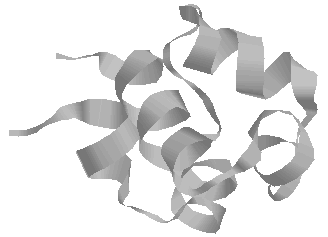
GOTO top
DNA structure
Here are two representations, again from the
same view point of a double helical DNA structure drawn as
balls and sticks:
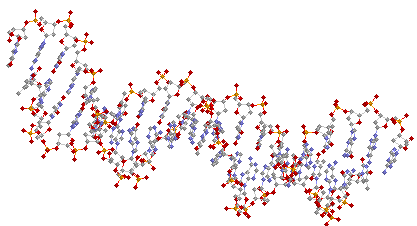
.... and as a cartoon:
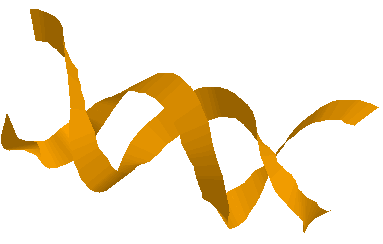
... and here is the space filling model of that DNA:
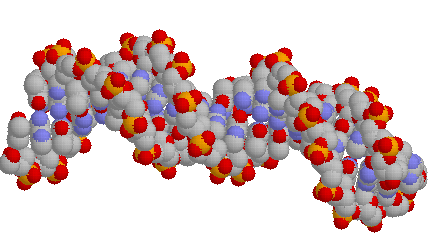
GOTO top
6: Genomes, gene regulation and protein biosynthesis
| Contents |
Genes and gene expression
Sequence relationships
|
| Abstract |
We explain the words/phrases, gene, chromosome, genome,
genetic system, haploid, diploid and allele. The expression
of many genes involves transcription (formation of RNA) and
translation (formation of protein) and there are significant
differences in these processes in bacteria and eukaryotes.
The relationship between DNA/RNA sequences is that of complementarity.
The relationship between an RNA sequence and a protein sequence is
the Genetic Code. In order to use this Code, it is important to
know which reading frame is in use.
|
| Objectives |
At the end of this topic you should have an understanding
of genes, genomes and gene expression including the
relationships between DNA, RNA and protein sequences.
|
| Why is this topic important? |
Much molecular biological data is derived from a knowledge
of DNA sequences.
|
| Why is it interesting? |
This a biochemical equivalent of language processing. A gene
sequence in DNA, in some sense, "means" the product(s)
of gene expression.
|
Genes and gene expression
I avoid giving strict definitions of the following words as many
derive from the period before the discovery of the role of DNA.
- gene a sequence of DNA (RNA in some viruses) which is
responsible for an inherited character. In many cases the gene
product is a protein
- chromosome a DNA sequence that exists as a continuous
molecule and contains several genes. Bacteria contain one
chromosome; other forms of life contain several (or many in the
case of ferns)
- genome all the chromosomes
- genetic system In animals and fungi, the mitochondria
have their own chromosomes the phrase "mitochondrial
genetic system", in contrast to the nuclear-cytoplamic
genetic system, is used to describe this. This is true also of
plants and they also have a chloroplast genetic system. Mitochonrial
and chloroplast genes are inherited maternally (also called
"non-Mendelian inheritance").
- haploid and diploid A cell that contains one
set of chromosome is haploid; one that contains two sets is diploid.
Bacteria are haploid. The somatic cells in tissues such as fingers,
heart, root, leaves... are diploid but germ cells are haploid. Some
fungi have haploid somatic cells. The word ploidy is
useful to describe organisms whose somatic cells contain other than
one or two sets of chromosomes (wheat is an example).
-
allele a version of a gene. Although this is somewhat
over-simplified, human eye colour, at least in European races, can
be regarded as having blue and brown alleles. This introduces the
idea that brown is dominant in this case as a person with
both alleles will have brown eyes. The same idea can be expressed
by saying that blue eyes are a recessive character.
Now we look at a picture that introduces some other words. The
expression of genes involves the production of messenger RNA (mRNA)
"RNA" in the picture.
In bacteria, genes for related functions are often clustered into
an operon. Such genes are cistrons and the RNA is
polycistronic mRNA. Eukaryotic genes frequently comprise
introns and exons: the intron sequences are
excised ("spliced") from the mRNA. The picture also
emphasises that there are start and stop sequences in both DNA and RNA
and, in the case of DNA, these are termed promoters and
terminators and that the expression of genes is commonly
regulated by transcription factors, usually proteins,
interacting with the DNA.
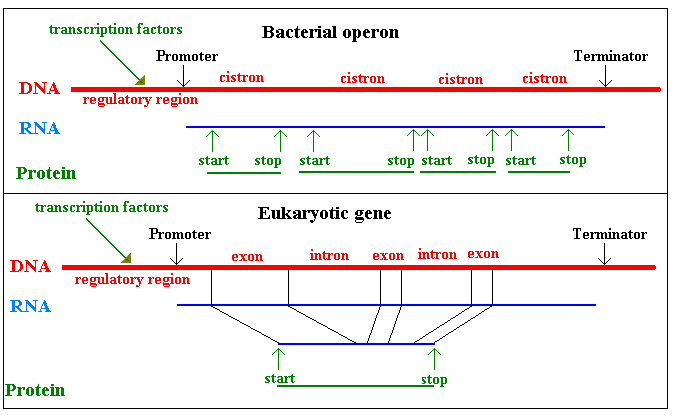
DNA acts a template for its own biosynthesis and for the
biosynthesis of RNA. RNA (more specifically one class of RNA)
acts a template for protein biosynthesis. In somewhat more detail:
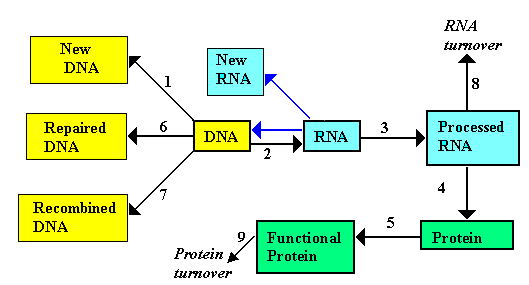
In this diagram the two processes shown with blue arrows and
without numbers occur only in cells infected with certain
types of virus. For the rest, the macromolecular components of
facets of the processes are summarised below. The numbers 1
to 9 refer to the diagram.
| |
Process names |
Enzymes |
Other components |
| 1 | DNA replication | protein |
protein, RNA |
|---|
| 1 | Regulation | protein |
DNA, RNA |
|---|
| 1 | Error correction | protein |
DNA, protein |
|---|
| 2 | Transcription | protein |
protein |
|---|
| 2 | Regulation | protein |
DNA, RNA |
|---|
| 3 | RNA modification and editing | protein, RNA |
protein, RNA |
|---|
| 4 | Translation | protein, RNA |
RNA |
|---|
| 4 | Regulation | protein, RNA |
protein, RNA |
|---|
| 4 | Error Correction | protein |
protein, RNA |
|---|
| 5 | Protein folding and modification |
protein | protein |
|---|
| 6 | DNA repair | protein |
protein, DNA |
|---|
| 7 | Genetic recombination | protein |
protein, DNA |
|---|
| 8 | RNA turnover | protein | protein, RNA |
|---|
| 9 | Protein turnover | protein | protein |
|---|
Several important points arise from these summaries
- Not all enzymes are proteins: some, the ribozymes, are
RNA molecules
- Errors that occur during DNA replication are corrected
- There are mechanisms (quite complicated in some cases) for
regulating macromolecular biosynthesis
- Macromolecules are not merely sysnthesised: RNA and protein
are turned over
- Many of the processes are thermodynamically highly unfavourable
- There is an important difference between the eukaryotic
system and that in bacteria, mitochondria and chloroplasts. Most
DNA in eukaryotes is in a separate cell compartment, the nucleus, and
RNA is transported to the cytoplasm for translation. In bacteria etc.
there is no nuclear membrane and not only do transcription and
translation occur in the same cell compartment but they can be
coupled, i.e. translation occurs while transcription is taking place.
Sequence relationships; the Genetic Code
DNA is a double stranded molecule; following transcription of a
gene mRNA is formed and translation of this leads to the
protein. These sequences are shown below in the case of the
gene for pCro. The transcribing strand of DNA is the template for
mRNA synthesis. In the picture, transcription and translation are
both from left to right. The non-transcribing (or plus) strand of
DNA has the same sequence as the RNA (if we replace the T's by U's)
and the molecular biological literature and databases usually just
show this plus DNA strand. In the picture, only the start of the
minus and mRNA sequences are shown and the promoter and terminator
regions are not shown.
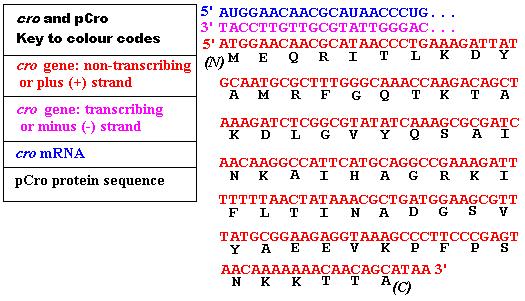
The relationship between the 2 DNA strands and between the transcribing
strand obey the rules of complementarity. The relationship between
the sequence of mRNA (or the + strand of DNA) and the protein is the
Genetic Code, which is a 3-letter non-overlapping code: 3 nucleotides
(usually referred to as "bases") constitute a codon
and, for example, the 2nd amino acid in pCro is E, the codon is GAA.
For a "biochemist's view" and "a modest suggestion
for a better view" click here.
Well, that seems fairly straightforward because "we knew"
how to relate the cro and pCro sequences. If we had known
that and merely had a piece of DNA sequence, there would be two
problems: (1) are we dealing with a plus or minus strand? and (2)
where do we start looking up the codons?
The example below is part of the cro gene.
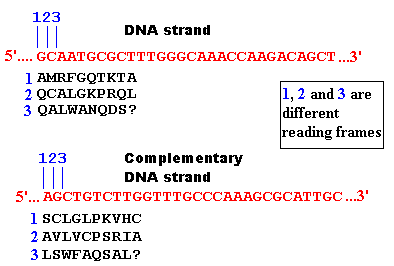
The complexities of the components
of the gene expression system ensure that that cells can solve the
problem. One task in bioinformatics is to enable us to do that.
GOTO top
7: Metabolic, regulatory and neural networks
| Contents |
Metabolism: some principles (i) Enzymes
Metabolism: some principles (ii) triphosphates and REDOX
Metabolic networks
Regulatory networks
Neural networks
|
| Abstract |
Enzyme catalysed reactions show characteristic saturation kinetics.
Metabolism is constrained by the requirement for overall favourable
(negative free energy change) processes. ATP can be regarded as
an "energy currency": the synthesis of ATP involves REDOX
reactions and, in the case of photosynthetic organisms the splitting
of H2O.
The biosynthesis of DNA, RNA and protein are examples of expensive
processes that have substantial demand on triphosphate utilisation.
"Metabolic pathways" (correctly metabolic nets) describe
the interconversions of metabolites. Regulatory networks can involve
many processes including the regulation of transcription and
translation. Although not a formal part of the module, some references
are given top the comparison between artificial neural nets and
the activities of neurons.
|
| Objectives |
At the end of this topic, you should understand the concept of
metabolism and the importance of free energy and metabolic and
regulatory networks.
|
| Why is this topic important? |
The subject is central to biological chemistry. More specifically
the accurate modelling of such networks is a major area of research
in contemporary bioinformatics.
|
| Why is it interesting? |
Biological networks resemble other types of nets in several respects but they
are characteristically robust and redundant.
|
GOTO top
Metabolism: some principles (i) Enzymes
Enzyme kinetics
The rates of simple enzyme catalysed reactions of the form
S → P1 + P2...
(we use S to stand for Substrate) follow the
Michaelis-Menten equation:
v = Vmax.[S]/([S]+KM)
where KM is the Michaelis constant and
Vmax is the maximum velocity. The
equation represents a hyperbola: -KM and
Vmax are asymptotes.
| ![Plot of v=Vm.[S]/(Km+[S])](mm.gif) |
Implications of the Michaelis-Menten equation:-
KM is charactersitic of the enzyme and
has the dimensions of
concentration (typical values would be of the order of 1 mM)
- KM is determined by the amount of enzyme in the
preparation used for the measurement
-
A small KM results in a steeper dependence of
velocity on [S] and this can be expressed that enzymes with low
KMs have high affinity for their substrate.
Limitations of this treatment:-
We have not considered reactions of the type
S1 +S2 → P1 + P2...
However there are such equations but they depend the mechanism
of the enzyme catalysed reactions -
More complicated kinetics are involved in allosteric enzymes
in whcih the binding of compound (which may or may not be a
substrate) affects the activity of the enzyme.
-
Equally we have left unexplored the properties of enzyme
inhibitors but note here that competitive inhibitors
reduce KM but leave Vmax
unchanged.
-
The major limitation to this type of equation is that
KM values are derived from rate constants
describing the association and dissociation of substrate(s) with
enzyme and the dissocation of product(s). Biochemists express this
by stating that these are steady state kinetics and note
gloomily that many enzyme catalysed reactions in vivo are
not at the steady state.
Nevertheless we can learn:-
Enzyme catalysed reactions are saturable which gives us
in principle a simple chemical mechanism for recognising thresholds
and performing a switch.
-
Kinetics of the Michaelis-Menten type apply equally to transport
processes. To take two examples:
- the reason why plants can
scavange incredibly low concentrations of soluble phosphate from
soil is that their roots (or in some cases symbioitic fungi)
have phosphate uptake systems with very low apparent
KMs
-
imagine you are resting after a meal of fish and chips, cream
cakes and sweet tea. Your blood glucose will be very high and
glucose will enter your red blood cells but not in a great rush.
Glucose transport in this case is thermodynamically favourable but
it does not diffuse into your blood by simple dialysis. It enters through
a process of facilitated diffusion whose rate is
determined by a red-cell glucose transport protein.
GOTO top
Metabolism: some principles (ii) triphosphates and REDOX
ATP
5'-Adenosine triphosphate, ATP, is sometimes called the "energy
currency" of the living cell. This is because its hydrolysis and
alcoholysis are very favourable (large -ve free energy change)
reactions. In the picture we draw attention to phosphate ester
bonds (blue) and phosphoanhydride bonds
(red). These latter are sometimes unhappily
referred to a "high energy bonds". The sketch
also introduces some further abbreviations including Pi
for inorganic phosphate and PPi for pyrophosphate.
Note that PPi is a phosphoanhydride.
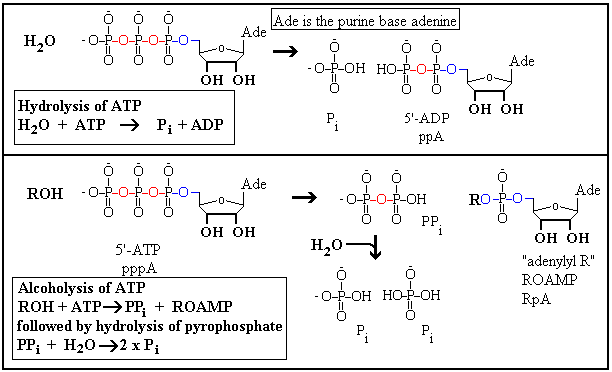
The importance of ATP and the other nucleoside triphosphates is that
their reactions can supply the necessary free energy change to drive
forward otherwise thermodynamically extremely unfavourable processes.
We shall exemplify this with nucleic acid and protein synthesis but
before tackling that, what processes allow the sysnthesis of the
very reactive phosphoanhydride bonds? The answer is that the
formation of phosphoanhydrides is coupled to very favourable processes
and, in the case of an aerobic organism living on fish, chips and
cream cakes, there are two sources of ATP. The breakdown of larger
molecules of carbohydrate and lipid (fat) leads to some ATP synthesis
itself but the larger source is in respiration. Sugars and fats are
chemically reduced and they themselves can become oxidised and
concommitantly reduce a specialised nucleotide called NAD (nicotinamide
adenine dinucleotide). NAD and its reduced form can be represented
by NAD+ and NADH/H+. Consider the following
reaction: the only background information needed is that lacate
can be oxidised to pyruvate. Here is the reaction:
pyruvate + NADH/H+
↔
lactate + NAD+
That is an example of a REduction/OXidation (REDOX) process.
To generate ATP (i.e. to form the phosphoanhdride bonds),
NADH/H+ provides via a pathway called the electron
transport chain the reducing power to reduce O2 to
H2O. Certain steps in the chain are thermodynamically
so favourable that they are coupled to the generation of ATP.
The pathway that generates the NADH/H+
in the first place results in decarboxylation of carboxylic acids and
thus we have the products of human respiration: H2O and
CO2.
Photosynthetic organisms, plants and cyanobacteria, harvest radiation
from sunlight and both fix "CO2" (i.e. reduce
it to form sugars) and generate O2 by splitting
H2O.
Now come to my promised examples of the use of ATP and other
triphosphates in the thermodynamically difficult jobs of making
polynucleotides and polypeptides.
For the polynucleotides we shall consider the chain extension
reactions and use a slightly expanded version of the abbreviated
sequence. We shall draw the 3'-terminus of a polynucleotide as
....NpNpNpNOH
, i.e. N for nucleosides and
a small OH as the 3'-terminal OH. Further we draw the template strand
(always DNA) in black, the growing strand in
blue and the next residue in
red. The enzymes (RNA- or DNA- polymerases)
are only parts of the machinery of DNA replication and transcription.
3'...ApApCpTpGpGpCpApApCpTpApApGpGp...5'
5'...TpTpGpApCpCOH
+
pppGOH →
3'...ApApCpTpGpGpCpApApCpTpApApGpGp...5'
5'...TpTpGpApCpCpGOH
+ pp
pp →
p + p
These two favourable reactions allow the unfavourable formation
of a new phosphodiester link and the unfavourable entropy change
involved in selecting just one (G in this case) out of a possible
of 4 nucleoside triphosphates.
With protein synthesis there are two separate steps: activation of
amino acids and peptide bond formation. An essential component of
the system is transfer RNA (tRNA) and we shall represent its C-terminus
as above. Each amino acid has its own small family of tRNAs. The
activation step takes place in two stages. We represent an
amino acid thus H2NaaCO2H and ATP as pppA:
(1) H2NaaCO2H + pppA
→ H2NaaCOpA + pp
(2) tRNA...pAOH + H2NaaCOpA → H2NaaCOOAp...tRNA + pA
|
H2NaaCOOAp...tRNA is aminoacyl tRNA,
henceforth H2Naa-tRNA
remembering that the - is an ester
bond.
Each amino acid has not only its own (family of) tRNA(s) but also
its own aminoacyl tRNA synthase enzyme that catalyses
both of the above reactions.
Messenger RNA (mRNA) is the template for translation. Ribosomes
are ribonucleoprotein particles involved in the translation process.
They can be regarded as molecular motors with associated enzyme
acitivity with a mechanical decoding
machinery. Essentially the ribosome reads mRNA like a tape in
an old fashioned computer and the processes (again we ignore the
problems of starting and starting and concentrate on the elongation
process) can be regarded as consisting of a few stages:
- there is a growing chain of polypeptidyl-tRNA attached
to a suitable site and an aminoacyl-tRNA is selected by nucleic
acid complementarity between the codon (in mRNA) and a sequence,
the anticodon in the aminoacyl-tRNA. This process involves
proteins called elongation factors and one GTP molecule is
concommitantly hydrolysed to GTP + Pi.
- The polypeptidyl tRNA and
aminoacyl tRNA react:
(N)....aaX-tRNAX
+
H2NaaY-tRNAY
→
(N)....aaXaaY-tRNAY +
tRNAX
This reaction is catalysed by a ribozyme,
peptidyl transferase, a component of the ribosome and does
not require hydrolysis of ATP or GTP. -
The mRNA "tape" has to be threaded through the ribosome and,
following conformational changes, a new site for aminoacyl-tRNA
is brought into register over the next codon. This process is
translocation and requires hydrolysis of GTP.
GOTO TOP
Metabolic networks
Well, my erstwhile colleagues might comment that I'm trying to condense the
entire subject of biochemistry into a few lines of "prose"
but here is an attempt (4 points):
- Biochemical pathways are actually components of networks
- Thermodynamic considerations dominate the subject
- There are 2 useful nouns
-
catabolism (adjective catabolic) metabolism that results in the
breakdown of more to less complex substances, i.e. -ve free energy changes
- anabolism (adjective anabolic) metabolism that results in the
biosynthesis of more complex substances, i.e. +ve free energy changes
- The networks are described as
"metabolic maps"*
and there are several resources on the Web, e.g.
*They are graphs rather than maps but the
metaphor might have been be derived from "maps" such as that of the London Underground.
GOTO top
Regulatory networks
Regulation can involve several chemical interactions:
Rearrangment of DNA sequences
Alternative splicing of mRNA precursors
Interaction of transcription factors and DNA
- the transciption factors can repress transcription
- this can require binding of a co-repressor to the repressor
- this can be inhibited by the binding of an inducer to the repressor
- very "simple" mechanism: is the repressor present or absent?
- the transciption factors can activate transcription
Translational control
- Attenuation
- masking of translational iniation
- by antisense RNA
- by RNA-protein interactions
Enzyme regulation-
substrate availability, saturation kinetics etc.
- allosteric mechanisms
This list is by no means comprehensive: very approximately we can
regard the list from 1 to 5 as representing coarse to
fine control. Not all mechanisms apply to all cells and all
circumstances and it must be emphasised that these mechanisms apply
to components of what are typically extensive regulatory networks.
These are just a few generalisations and examples.
Examples of 1 and 2 can be found in the immune system
and some of the methods used by invading micro-organisms to evade it.
Another example of 1 is the ability of yeasts to undergo a
"sex change", correctly a switch in mating type.
We return shortly to 3 but note here that we have left
a lot out of 3.2 and examples of this include many processes
controlled by hormones. 4.1 involves the optional recognition
or non-recognition of transcriptional terminators: the mechanism relies
on the fact that RNA polymerase, recognises a terminator sequence in
its transcript, not in its template. 4.2.1 is perhaps worth
considering briefly: the method is used in the genetic engineering of
certain crop plants and as an approach to treatment of genetic
diseases but it does occur naturally, for example in E. coli.
Again, a gross over-generalisation: 5 is very characteristic
of cells in animals, plants and fungi but bacteria tend to rely on
3.1, 3.2, 4.1 and 4.2 for regulating the
activities of enzymes because they are designed to respond to changes
in their ecoment but there is simply not
enough room in (for example) an E. coli cell for biochemically
significant amounts of every enzyme that might come in useful.
I shall not consider these regulatory networks in detail but rather shall
just show one example of how a protein considered previously can
effect a switch: an irreversible one in this case. cro is
a gene in lambdoid phages. Phages (short for bacteriophages)
are bacterial viruses. Some, including the lambdoid phages, are said
to be temperate which means that, following infection, the
phage can either be reproduced with (in the case of these phages)
lysis of the cell or can establish a state known as
lysogeny in which the phage DNA is incorporated into the
host's genome as a prophage. (This use of the affix pro
is used more generally: e.g. HIV forms a provirus in infected
cells.) The sketch below shows part of a lambdoid prophage.
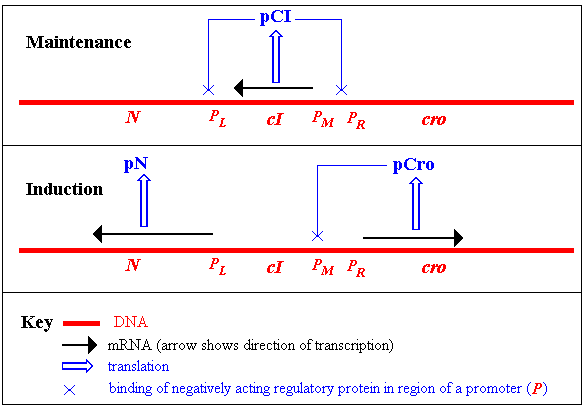
In a lysogen, the only prophage gene to be expressed is cI
but if pCI is inactivated, pCro appears in the cells and, following
this, the first stage in induction, cI is never
expressed again: the E. coli cell is doomed and lysis and
release of more phage particles is sure to occur after about 3/4 of
an hour. The system is designed to allow prophage to be propagated as
part of the E. coli genome until the cell is at risk: a
system, not described here, ensures that pCI is destroyed if the
cell is exposed to UV or ionising radiation or to genotoxic chemicals.
Given that, the phage make use of their 50 genes to construct new
phage particles, to traverse the environment and find a new host.
Two final points about the cI-and cro (also known as
immunity) system.
- It is vastly over-simplified in the sketch: there are 3 binding
sites for pCI and pCro in the regions of
(i) PL and (ii) the overlapping
PR and PM sites and pCI and
pCro actually compete with each other bu titrating out the more
favourable sites.
-
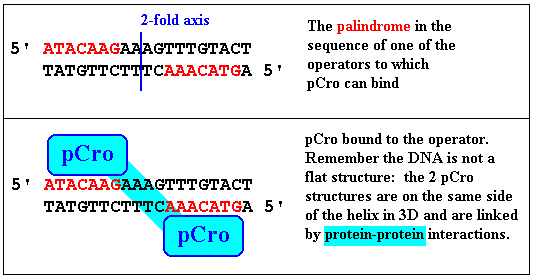
The sites these proteins bind to are operators and are
palindromes. The binding of pCro to one of its palindromic
sites in DNA (such regulatory DNA sequences for -vely acting regulatory
proteins are, in bacteria, referred to as operators. Two
pCro molecules and the operator form a ternary complex.
GOTO top
Neural networks
This section is included to give computer
scientists interersted in artificial neural nets (ANNs) access to
intructory material on real neural nets. One type of simple
ANN devised for machine learning might rely on a sigmoidal
or logisitic activation function:
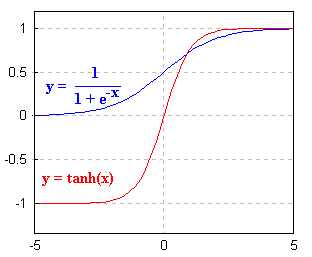
... and might be used in a "feed-forward"
ANN for machine learning;
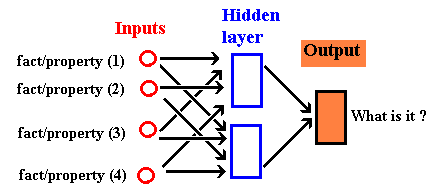
Of course there are many different designs for
ANNs and in particular we refer to the completely connected Hopfield
networks but my task here is to draw attention to differences between
neurons in vivo and the artificial neurons, perceptrons...
("units" in the diagram).
First there are more neurons (1011 in a human brain).
Secondly, they do not look much like computer hardware or software:
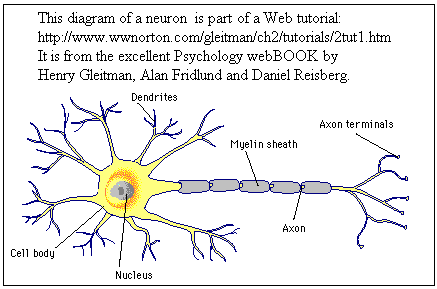
Thirdly the anatomy of the
brain involves the very precise organisation of neurons, e.g. in
layers achieved by having axons of different lengths (the axons are
electrical conductors and the myelin is an insulator).
The diagram highlights them.
Here is
another more anaimated) diagram and one from a more
psychological perspective.
Link 1 and
Link 2 to tutorials on computational aspects.
Oct. 2009 replacement of defunct links.
GOTO top
8: Classification and ontologies in biological sciences
| Contents |
Speciation
Enzymes and Molecules |
| Abstract |
Biological classification or taxonomy was originally based on the
scoring of characters in specimens. The systematic method of doing
this was an early example of computer aided linkage analysis.
Modern taxonomies rely on macromolecular sequences. Enzymes are
classified in a system of "EC numbers". The heirarchy in
this case is less easy to parse.
|
| Objectives |
At the end of this topic, the student should have a working knowledge
of the principles and practice of biological taxonomy and to be
familar with resources to relate EC numbers and enzyme activities.
|
| Why is this topic important? |
Classification is important in relating current research studies to
older literature descriptions.
|
| Why is it interesting? |
Biological taxonomy is an example of an ontology.
|
GOTO top
Speciation
The biological description and classification of organisms dates
from a period of natural history. Most people are familiar with
the idea that humans belong to the species Homo sapiens and
therefore to the genus Homo of which H. sapiens is the
only surviving species. We also have a sense that there are larger
groupings: e.g. H. sapiens and monkeys are part of a tighter
grouping than, for example, H. sapiens and rabbits and all of
these are more "related" to one another than they
are to fish, mushrooms, cauliflowers, seaweed, bacteria. The general
name for categories such as species, genera, families... is
taxon. The first systematic approach to defining taxa was
made by Adanson in the XVIII century. The idea recognised that there
are variations in populations of, e.g. H. sapiens but if we
were to score characters (true or false) for enough specimens, the
specimens could be put into taxa. A simple, not to say rather
ridiculous, example follows.
| ↓ Character
| Specimen → |
1 | 2 | 3 | 4 | 5 | 6 | 7 |
8 | 9 |
|---|
| has 4 legs, 1 at each corner |
1 | 1 | 1 | 1 | 0 | 1 | 1 |
1 | 1 |
| has a tail |
1 | 1 | 1 | 1 | 0 | 1 | 1 |
0 | 1 |
| has course hair |
0 | 0 | 1 | 0 | 0 | 1 | 0 |
0 | 1 |
| has soft fur |
1 | 1 | 0 | 0 | 0 | 0 | 1 |
1 | 0 |
| makes miaowing sound when stroked |
1 | 0 | 0 | 0 | 0 | 0 | 1 |
1 | 0 |
| is coloured black |
0 | 1 | 0 | 0 | 0 | 0 | 0 |
1 | 1 |
| makes barking noise when needs feeding |
0 | 0 | 1 | 0 | 0 | 1 | 0 |
0 | 0 |
From this we may deduce that specimens 1, 2, 7 and 8 may be cats
(although 2 may have a sore throat and 8 presumably comes from the
Isle of Man) and 3, 6, 7 and 9 may be dogs (of which 7 and 9 are
laudably quiet). We note that being black is a variable character
in both groups and that we cannot say much about specimen 5.
Nevertheless Adanson was well ahead of his time: this is an
objective method and, given hundreds or thousands of specimens and
characters there is a system of classification which worked very
effectively for many years, notably in the characterisation of
bacteria. Of course it awaited advances in statistics and the advent
of computers to construct the clusters and to produce hierarchic taxa.
Today the clustering might well be done by K-means but computational
methods of linkage analysis were pioneered by Sneath and Sokal for
Adansonian analysis. In theory at least the grouping of organisms
into larger and larger clusters, e.g. species, genus, family, order,
class... can be achieved by such methods. Modern taxonomies are based
on clustering of macromolecular sequence data rather than
characters such as colour etc. but, rather remarkably, the fit between
classical and such "molecular" taxonomies is quite good
provided we do not rely on "classical" methods to
investigate relationships between deeply divided taxa.
There are several problems surrounding these taxa of which
here are a few.
| | Question | Possible answer | Problem |
|---|
| 1 | What is a species? |
defined by saying (i) 2 members of a species can interbreed
but (ii)
members of different species cannot |
Any gardener or keeper of pet fish can tell you (ii)
is not true and (i) fails for organisms that do not reproduce
sexually. |
|---|
| 2 | How are members of a taxon related? |
They had a common ancestor |
Among the bacteria (possibly higher organisms) genes or clusters
of genes can be transferred between very unrelated organisms. |
|---|
| 3 | How do we decide how the boundaries of a taxon
might be defined? |
let's say that a certain degree of similarity, sequence
homology etc. defines a species etc.: S% for a species, G% for a genus,
F% for a family... where S<G<F ... |
Well that's not going to work too well! Using the criteria of
bacterial taxonomy, humans, chimpanzees and gorillas would all be
minor variations within the same species and yet they are not
even in the same genus. |
|---|
Molecular biology and bioinformatics have resolved some of the
problems of biological taxonomy. Sequence analysis has enabled
the objective classification of organisms and consequently the
familiar phylogenetic trees of biology text books can be
extended considerably. The nature of the macromolecule to sequence
depends on the nature of the problem we set ourselves. In the sketch
I show 3 phylogenetic trees. Although the scales are not given,
the qualitative conclusions are reasonably acceptable.
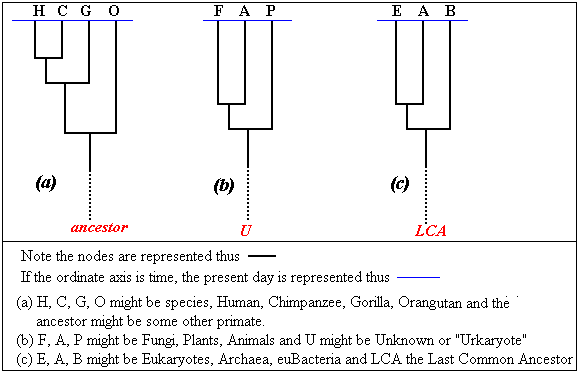
If we are prepared for a root node, these are binary trees. In the
case of 4 great apes, the root could be an animal believed to be
an ancestor (e.g. a monkey as suggested in the sketch) or a fish.
Such a root is referred to as and outgroup. With the
three major divisions of life (b), we do not know what the ancestor
might be as there is neither fossil record nor an extant intermediate
form to help us but the word urkaryote has been coined to
describe some eukaryotic antecedent with mitochondrial or
chloroplast genetic systems. Part (c) describes to the current
version of the history of life. Here we can estimate the
time axis: some 4x109 years. The Archaea resemble
Eubacteria in that they have no nuclear envelope and are hence
prokaryotes. Examples of archaea include the purple
bacteria that grow in saturated saline solution and in several other
extreme environments. Such trees can be constructed by macromolecular
sequence comparison. In order to construct a tree such as (c), we need
macromolecules which are present in forms of life, fulfil the same
role in all of these and evolve slowly: rRNAs are the molecules of
choice and it was analysis of the sequences of these that led to the
recognition of the archaea (formerly "archaebacteria") by
Carl Woese. Subsequenctly several other features link the archaea and
the eukaryotes, notably the presence of introns. For (b), rRNA
sequence data are reinforced by RNA polymerase sequence comparisons
and certain metabolic similarities. The great ape classification
relies on several points such as the sequence relatiobships of a
a characteristic and variable region of mitochondrial DNA (mtDNA):
the short sequence is not required for any known mitochondrial
function and thus there is no (or at least little) selection
against mutation. Moreover, the role of mitochondria in respiration
results on a relatively high concentration of DNA-reactive reactive
oxygen species (ROSs: examples include the superoxide ion
-O2 and its breakdown products).
GOTO top
Enzymes and molecules
Biochemists established several years ago a classification for
enzyme-catalysed reactions. The classification is in the hands of
a committee, the Enzyme Commission, EC, which is perhaps a
bioichemical counterpart of IEEE or ISO. In considering the EC
systme it is important to recognise that it is not a
classification of enzymes as proteins but rather of the reactions
classified so, for example two enzymes might be structurally similar
but catalyse different reactions. EC classifications are known as
"EC numbers" and take the form EC N1.N2.N3.N4. There are
plenty of W3 sites that summarise EC numbers but
this
is an official one. In summary there are six values of N1 from 1-6
and these plus a few extra examples are listed in the table below.
| N1 | Description | number of N2's |
examples |
|---|
| 1 | Oxidoreductases | 19+"97" |
|
|---|
| 2 | Transferases | 9 |
|
|---|
| 3 | Hydrolases | 13 |
|
|---|
| 4 | Lyases | 6+"99" |
4.1 carbon-carbon lyases
4.2 carbon-oxygen lyases |
|---|
| 5 | Isomerases | 5+"99" |
|
|---|
| 6 | Ligases | 5 |
6.1 forming carbon-oxygen bonds
6.4 forming carbon-carbon bonds |
|---|
A full EC number would be, for example, EC 1.1.1.1
(alcohol dehydrogenase). Three points deserve making as they are
important in any software that might be used in parsing EC numbers.
- as a tree it is very unbalanced
- large numbers are used to refer to reactions awaiting
classification (e.g. the EC 1.97 enzymes)
- there are no rules for determining the nodes. For example,
the EC 4.1 enzymes break C-C bonds but EC 6.4
enzymes form them.
The EC system and biological taxonomy raise problems in bioinformatics
because they are, to some extent rigid, and do not respond readily
to advances in the molecular sciences. However they fulfill a separate
role. They are useful in defining organisms or enzymes that are
described in the literature before the current molecular understanding.
Accepting this caveat, analysis of biological phylogenetic trees and
the EC system are different problems. The fact that the EC system
is a general tree (or possibly a forest of 6 such trees) is trivial as
any general tree can be transformed into a binary tree. However,
analysis of biological classification is a tree traversal problem
whereas the EC system, despite its reliance on heirarchies, is
essantially a look-up table.
One of the successes of bioinformatics has been the developoment of methods for classifying macromolecules
on the basis of sequence and structure. An important generalisation is that structure is more highly
conserved than sequence.
GOTO top
9 Recombination, repair, rearrangment and evolution
| Contents |
Recombination (1)
Recombination (2)
DNA repair
Evolution
|
| Abstract |
Reciprocal recombination, not be confused with chromosome inheritance
occurs during meiosis. Site-specific recombination is important in
several events notably transposition of genes. Enzymes monitor DNA
for evidence of damage or erroneous replication and effect repair
frequently by re-synthesis. DNA biochemistry provides a mechanism
for biological evolution and site-specific recombination and allied
processes provide a mechanism for lateral gene transfer between
bacteria.
|
| Objectives |
At the end of this topic, the student should be familar with DNA
rearrangements in genetic recombination, the response of cells to
damaged DNA and should have an understanding of the role of
molecular biology in providing plausible mechanisms for biological
evolution.
|
| Why is this topic important? |
Bioinformatics relies extensively on principles of change and
evolution.
|
| Why is it interesting? |
This topic has generated one of the most powerful heuristics
in computer science: the genetic algorithm.
|
GOTO top
Recombination: part 1
Below we look at the result of "reciprocal recombination".
Blue and red
parental double stranded DNA molecules have crossed over at the
points marked X so that the sequences are changed. The sketch
omits both biological and biochemical details but we can point out that
the event occurs during the process of meiosis (or its
equivalent) and that intermdiates not shown here involve a
heteroduplex intermediate, i.e. a DNA molecule with one
blue and one red
strand. The numbers 1-9 represent genetic markers (let's
just oversimplify and say "genes") and if the
blue and red
alleles are phenotypically different we find
a new assortment of genes, e.g. the upper product of the cross
is
1 2 3 4 5
6 7 8 9
The phenotype is
the property. For example let us imagine that the cross occurs in
some kind of fish with coloured spots and fins, that gene
2 determines the spot
colour (blue or
red) and that gene 4 determines the
fin colour. The parental phenotypes are
blue fins, blue spots and
red fins, red spots. We have below the
genotype of an offspring that has red fins
(4) and blue
spots(2).
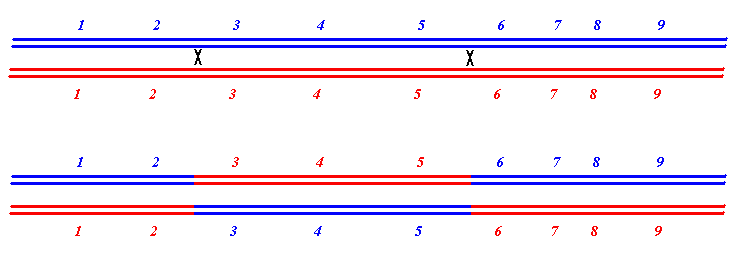
Before we proceed, it is imortant not to confuse recombination of this
sort with the assortment of chromosomes first described by Mendel in
peas. The next cartoon shows some fish and we introduce a new
character. Their eyes can be either dark or yellow:
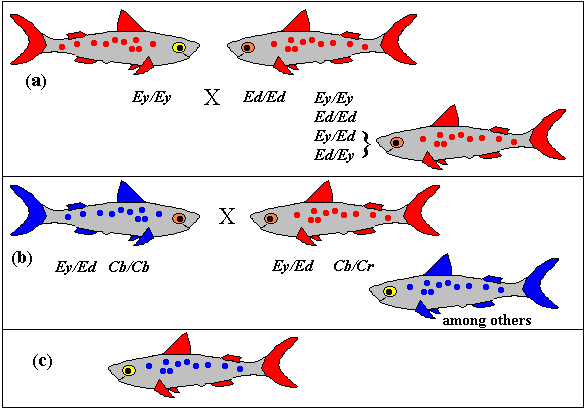
In (a) each parent has identical alleles for the eye colour gene
(Ey or Ed for yellow or dark eyes. Each offspring
inherits half of its chromosomes from each parent and the result in
this fictitious case is that 3/4 will have the dark eyed phenotype
because the Ed allele is dominant. In (b) we are
looking at the assortment of characters involving also fin-and-spot
colour. Note from the appearence of the right hand parent, the
red "allele" (Cr) is seemingly dominant. It is
possible to follow Mendel and calculate the expected frequencies of
phenotypes in the offspring. Fish (c) can only arise from crosses
such as that in (b) if recombination arises. Thus the C
gene is actually 2 genes (we have called them 2 and 4)
and offspring such as (c) will arise rarely. The genes 1 to
9 are said to be linked and chromosomes are thus examples of
linkage groups.
GOTO top
Recombination: part 2
The integration of the DNA of a lambdoid temperate
phage into a
bacterial chromosome is shown in the
next sketch. Note that the DNA is circularised, the integration
involves a single cross-over and, importantly, the gene order in
the phage and prophage DNAs are different.
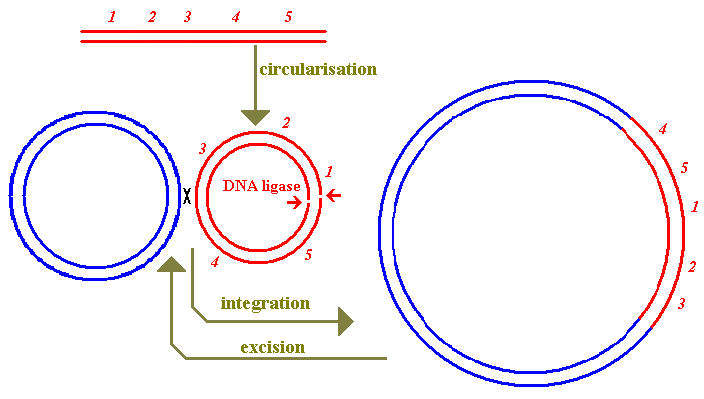
As the integration occurs at one of a very limited number of sites
(only one in the case of the E. coli chromosome and the
lambdoid phages) and a specific sequence in the phage DNA, this is
an example of site-specific recombination. Finally we
look at other examples (both taken from bacterial genetics) of
comparable processes. To simplify the diagrams, double stranded
DNA is drawn as a single line.
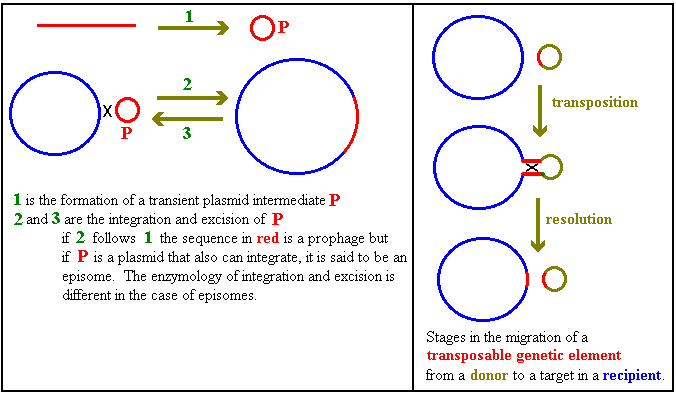
The left hand panel shows that the integration of prophage DNA has its
counterpart with certain plasmids ("episomes;")
that can integrate into the bacterial chromosome. In these cases the
"specific sites" are insertion sequences (ISs) and
as these are mobile genetic elements, episomes can integrate at a
wide variety of sites. The right hand side of the picture illustrates
one type of transposition in which a
transposable genetic element is copied from one site to
another. The cartoon is meant to show such an element being copied
from a plasmid to a bacterial chromosome but this is not the only
circumstance under which this can happen. The events we have just
covered have great implications for evolution but
here we just tidy up a few points of nomenclature and other
implications.
1. Transposable genetic elements that carry genes for functions other
than their own transposition are transposons
2. ISs are examples of transposable genetic elements
3. Transposons are not unique to bacteria: they cause genetic
instability in several organisms and were first discovered as a
cause for this in maize.
4. Certain bacterial transposons include genes for resistance to
anitbiotics, drugs and disinfectants.
5. Certain bacterial plasmids are conjugative which means
they can be "transferred" (actually they are copied as
DNA replciation is involved) from donor to recipient cell and
these cells to not need to belong to the same or even a related
species.
6. Conjugative plasmids are not the vehicles for gene transfer between
bacteria: certain phages can do this and phage-mediated gene transfer
from cell to cell is called transduction.
7. The importance of points 4-6 to the clinical management of
infectious diseases can hardly be overestimated. New arrangements
of drug-resistance genes are constructed by these mechanisms.
GOTO top
DNA repair
There is a lecture on this subject under Mol. Biol.
An important consequence of DNA repair is that it can
result in errors and hence in mutation.
Evolution
The basis of Darwinism* is
that variations arise in populations and certain of these variants
survive because they are in some sense fit. Darwin was unaware of
Mendel's pioneering study of genetics and of course neither had any
knowledge of molecular biology.
* I have avoided referring
to this as a "theory of evolution" because as a theory or
hypothesis it is ill formed: as far as we know it has only happened
once and cannot be tested. Perhaps Darwinism should be described as
a diagnosis. A more formal
account would be to observe that evolution is an example of
emergent properties which takes us back to where we
began.
The cartoon below illustrates a present day view of Darwinism as
it applies to 10 species (numbered in hexadecimal!).
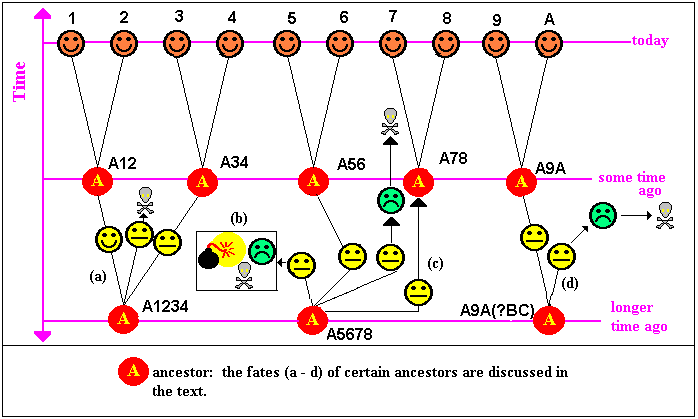
(a) Organisms 1-4 have a common ancestor A1234; we show one example of
variant that became extinct between A1234 and A12/A34.
(b) The fate of the lineage from A5678 is more complicated and we
conclude that one extinction was due to some kind of catastrophe.
(c) Some of A5678's decendents had a more successful history.
(d) Natural extinctions occurred early in the lineage from the
antecedents of A9A.
Evolution has proceeded at an uneven pace. One factor is that large
evolutionary change has a pre-requisite for DNA amplification and
the selective advantage may not increase fitness. DNA rearrangement
can result in big changes occurring over a short period of time
and changes in a very small number of genes can affect the
phenotype. For example, in recent human evolution, in which the
species Homo erectus was displaced more-or-less successively
by the H. sapiens types, Neanderthal, Cro-Magnon and Modern,
one of events may have been neoteny, the retention of
juvenile characteristics into adulthood and few mutations and/or
recombinations might in principle achieve this.
Lateral
gene transfer mediated by mechanisms described
earlier result in a more complicated pattern of
inheritance. Drug resistance and the migration of genes been organelles are ex\mples of this.
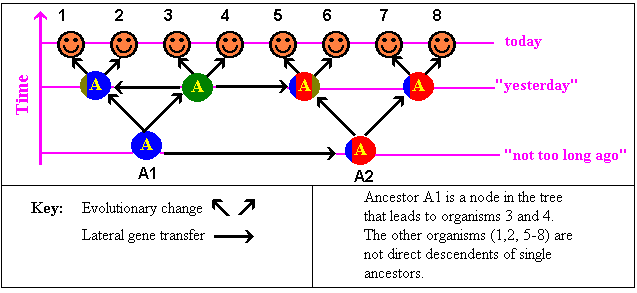
We have noted already the importance of the potentially very short
time scale involved in such rearrangements.
GOTO top
 Informatics describes the study and practice of creating, storing, finding, manipulating and sharing information. There are many other neologisms and phrases derived from this: bioinformatics, chemoinformatics, health informatics, nursing informatics, poli-informatics, for example.
Informatics describes the study and practice of creating, storing, finding, manipulating and sharing information. There are many other neologisms and phrases derived from this: bioinformatics, chemoinformatics, health informatics, nursing informatics, poli-informatics, for example.





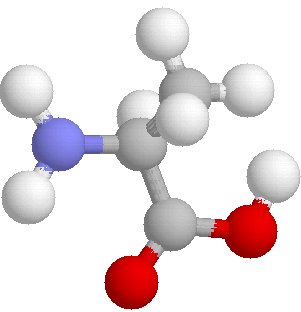
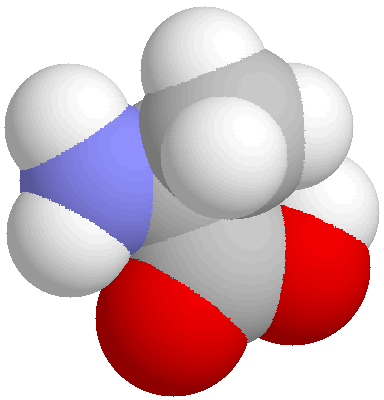


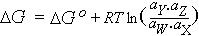
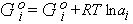
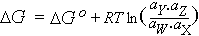






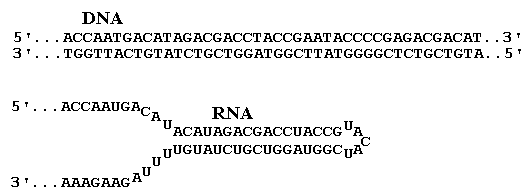


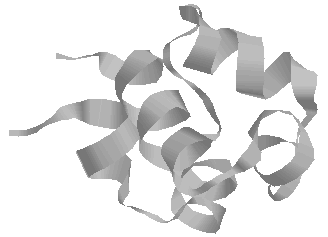

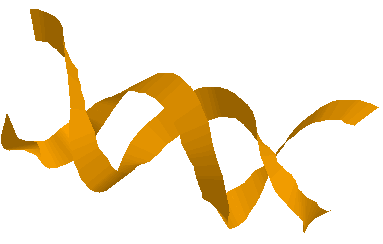





![Plot of v=Vm.[S]/(Km+[S])](mm.gif)












